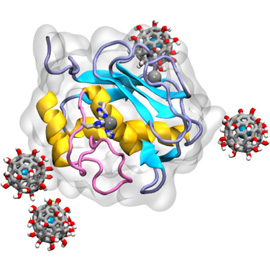
A computational model of an anticancer nanoparticle
September 7, 2012
Researchers have used computational modeling to precisely simulate how a drug inhibits a target enzyme known to spur cancer’s spread, capturing the interaction at the quantum-mechanical level, Technology Review reports.
They hope their work will uncover a way to specifically inhibit members of a whole class of cancer-linked proteins without causing as many side effects as existing drugs.
The authors showed that a buckyball molecule, which forms a cage enclosing a single atom of a heavy metal called gadolinium, can decrease the spread of pancreatic cancer in mice by inhibiting enzymes called MMPs, which help tumors reëngineer blood vessels to supply themselves with nutrients.
To capture the effects of the heavy metal ion on the nanoparticle and the enzyme, the researchers needed to examine the quantum mechanics of the interaction. That required a supercomputer — in this case, IBM’s Blue Gene.
With the computer’s help, the team identified the exact location where one of the enzymes, MMP-9, sticks to the nanoparticle. The model also predicted that the nanoparticle might clump together before interacting with MMP-9, and the authors were able to demonstrate that it does form clusters in aqueous solutions.
The study was published in an open-access article this week in the Proceedings of the National Academy of Sciences.