A hidden genetic code for better designer genes
October 1, 2013
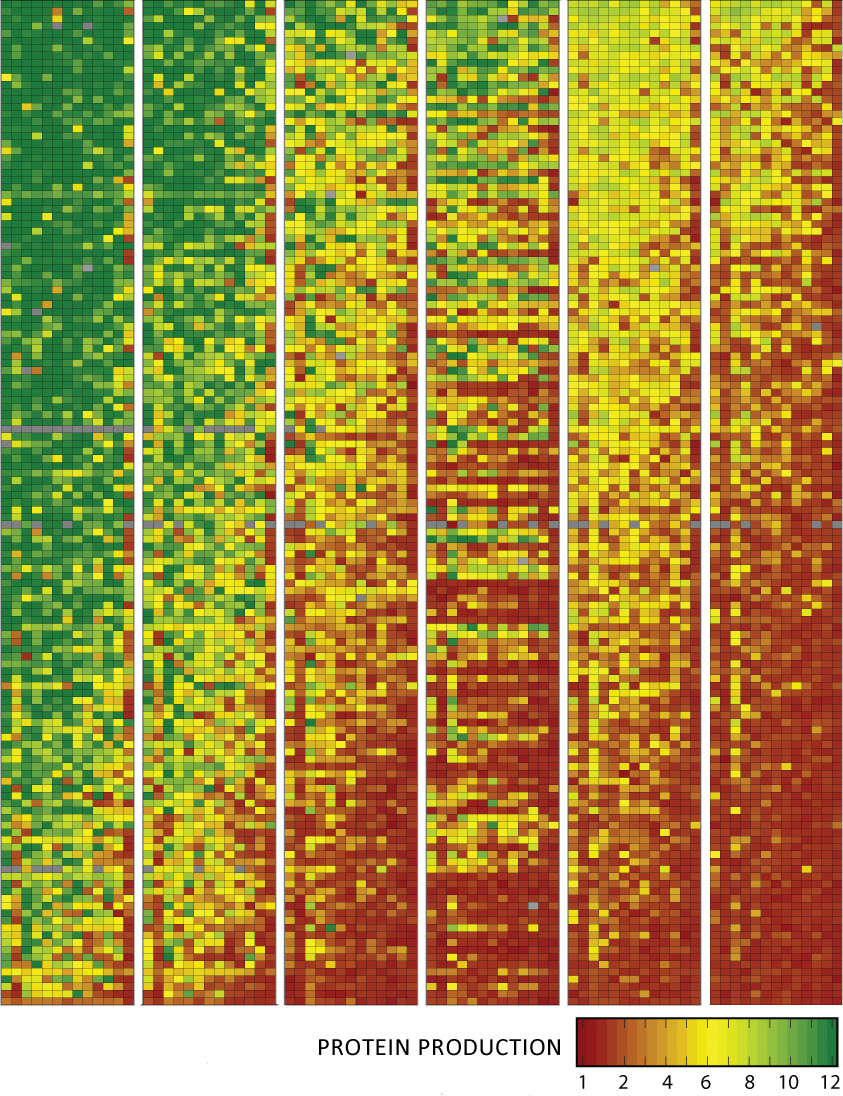
Wyss Institute scientists synthesized 14,000 snippets of DNA (individual pixels) and tested how well they promote protein synthesis. They found that those with rare ³words² (codons) near the start of bacterial genes made more protein (green pixels) by removing roadblocks to protein production. (Credit: Wyss Institute)
Scientists routinely seek to reprogram bacteria to produce proteins for drugs, biofuels and more, but they have struggled to get those bacteria to follow orders.
A hidden feature of the genetic code, it turns out, could achieve that. The feature controls how much of the desired protein bacteria produce, a team from the Wyss Institute for Biologically Inspired Engineering at Harvard University reported in the September 26 online issue of Science.
Rare condons
The researchers combined high-speed “next-generation” DNA sequencing and DNA synthesis technologies and found that using more rare words, or codons, near the start of a gene removes roadblocks to protein production.
The research team is headed by Sri Kosuri, Ph.D., a Wyss Institute staff scientist, George Church, Ph.D., a core faculty member at the Wyss Institute and professor of genetics at Harvard Medical School, and Daniel Goodman, a Wyss Institute graduate research fellow,
“Now that we understand how rare codons control gene expression, we can better predict how to synthesize genes that make enzymes, drugs, or whatever you want to make in a cell,” Kosuri said.
The findings could be a boon for biotechnologists,and could help synthetic biologists reprogram bacteria to make new drugs and biological devices.
To produce a protein, a cell must first make working copies of the gene encoding it. These copies, called messenger RNA (mRNA), consist of a specific string of words, or codons. Each codon represents one of the 20 different amino acids that cells use to assemble proteins. But since the cell uses 61 codons to represent 20 amino acids, many codons have synonyms that represent the same amino acid.
In bacteria, as in books, some words are used more often than others, and molecular biologists have noticed over the last few years that rare codons appear more frequently near the start of a gene. What’s more, genes whose opening sequences have more rare codons produce more protein than genes whose opening sequences do not.
No one knew for sure why rare codons had these effects, but many biologists suspected that they function as a highway on-ramp for ribosomes, the molecular machines that build proteins. According to this idea, called the codon ramp hypothesis, ribosomes wait on the on-ramp, then accelerate slowly along the mRNA highway, allowing the cell to make proteins with all deliberate speed.
But without the on-ramp, the ribosomes gun it down the mRNA highway, then collide like bumper cars, causing traffic accidents that slow protein production. Other biologists suspected rare codons acted via different mechanisms. These include mRNA folding, which could create roadblocks for ribosomes that block the highway and slow protein production.
A high-speed, multiplexed method
To see which ideas were correct, the three researchers used a high-speed, multiplexed method that they’d reported in August in The Proceedings of the National Academy of Sciences.
First, they tested how well rare codons activate genes by mass-producing 14,000 snippets of DNA with either common or rare codons; splicing them near the start of a gene that makes cells glow green, and inserting each of those hybrid genes into different bacteria. Then they grew those bugs, sorted them into bins based on how intensely they glowed, and sequenced the snippets to look for rare codons.
They found that genes that opened with rare codons consistently made more protein, and a single codon change could spur cells to make 60 times more protein.
“That’s a big deal for the cell, especially if you want to pump out a lot of the protein you’re making,” Goodman said.
The results were also consistent with the codon-ramp hypothesis, which predicts that rare codons themselves, rather than folded mRNA, slow protein production. But the researchers also found that the more mRNA folded, the less of the corresponding protein it produced — a result that undermined the hypothesis.
To put the hypothesis to a definitive test, the Wyss team made and tested more than 14,000 mRNAs — including some with rare codons that didn’t fold well, and others that folded well but had no rare codons. By quickly measuring protein production from each mRNA and analyzing the results statistically, they could separate the two effects.
The results showed clearly that RNA folding, not rare codons, controlled protein production, and that scientists can increase protein production by altering folding, Goodman said.
The new method could help resolve other thorny debates in molecular biology. “The combination of high-throughput synthesis and next-gen sequencing allows us to answer big, complicated questions that were previously impossible to tease apart,” Church said.
“These findings on codon use could help scientists engineer bacteria more precisely than ever before, which is tremendous in itself, and they provide a way to greatly increase the efficiency of microbial manufacturing, which could have huge commercial value as well,” said Wyss Institute Founding Director Don Ingber, M.D., Ph.D. “They also underscore the incredible value of the new automated technologies that have emerged from the Synthetic Biology Platform that George leads, which enable us to synthesize and analyze genes more rapidly than ever before.”
The work was funded by the Department of Energy, the Office of Naval Research, Agilent Technologies, an NSF Graduate Research Fellowship to Daniel Goodman, and the Wyss Institute.