Chemical imaging brings cancer tissue analysis into the digital age
January 31, 2014
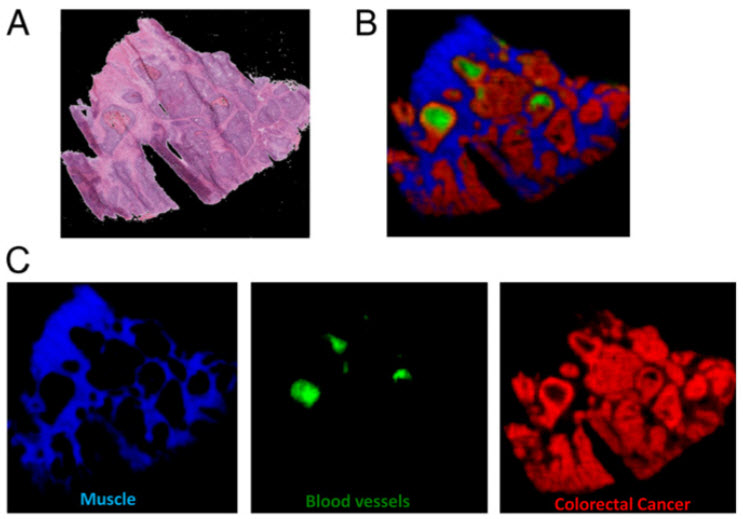
A section of bowel tissue as an optical image (A) and using mass spectrometry imaging and pattern recognition to identify tissue types (B and C) (credit: Imperial College London)
Imperial College London researchers have developed a new method for analyzing biological samples based on their chemical makeup that could transform the way medical scientists examine diseased tissue.
When tests are carried out on a patient’s tissue today, such as looking for cancer, the test has to be interpreted by a histology specialist, which can take weeks to get a full result.
Scientists have proposed using mass spectrometry imaging (MSI), which uses technologies that reveal how hundreds or thousands of chemical components are distributed in a tissue sample. But currently proposed MSI workflows are subject to several limitations, including nonoptimized raw data preprocessing, imprecise image coregistration, and limited pattern recognition capabilities.
In Proceedings of the National Academy of Sciences (open access), the Imperial College London researchers have now outlined a comprehensive new strategy for effectively processing MSI data and building a database of tissue types.
How new MSI process works
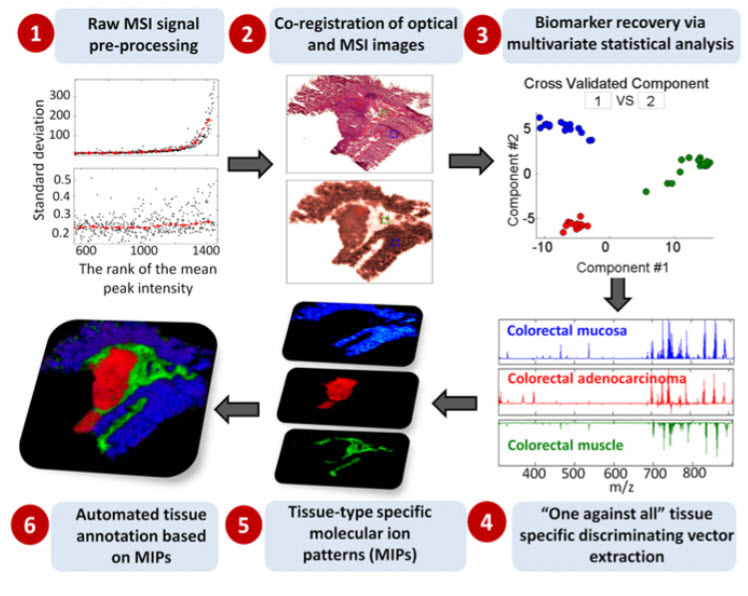
Overall computational workflow for exploration of region-specific
lipid biochemistry using MSI platforms (credit: Kirill A. Veselkov et al./PNAS)
In MSI, a beam moves across the surface of a sample, producing a pixelated image. Each pixel contains data on thousands of chemicals present in that part of the sample. By analyzing many samples and comparing them to the results of traditional histological analysis, a computer can learn to identify different types of tissue.
A single test taking a few hours can provide much more detailed information than standard histological tests, for example showing not just if a tissue is cancerous, what the type and sub-type of cancer, which can be important for choosing the best treatment. The technology can also be applied in research to offer new insights into cancer biology.
According to Kirill Veselkov, M.D., corresponding author of the study from the Department of Surgery and Cancer at Imperial College London, “MSI is an extremely promising technology, but the analysis required to provide information that doctors or scientists can interpret easily is very complex. This work overcomes some of the obstacles to translating MSI’s potential into the clinic. It’s the first step towards creating the next generation of fully automated histological analysis.”
“There have been relatively few major changes in the way we study tissue sample pathology since the late 19th century, when staining techniques were used to show tissue structure,” Professor Jeremy Nicholson, Head of the Department of Surgery and Cancer at Imperial College London, added. “Such staining methods are still the mainstay of hospital histopathology; they have become much more sophisticated but they are slow and expensive to do and require considerable expertise to interpret.
“Multivariate chemical imaging that can sense abnormal tissue chemistry in one clean sweep offers a transformative opportunity in terms of diagnostic range, speed and cost, which is likely to impact on future pathology services and to improve patient safety.”
The technology will also be useful in drug development. To study where a new drug is absorbed in the body, pharmaceutical scientists attach a radioactive label to the drug molecule, then look at where the radiation can be detected in a laboratory animal. If the label is detached when the drug is processed in the body, it is impossible to determine how and where the drug has been metabolized. MSI would allow researchers to look for the drug and any metabolic products in the body, without using radioactive labels.
The research was funded by Imperial College London’s Junior Research Fellowship scheme, awarded to Dr Kirill A. Veselkov; the National Institute for Health Research Imperial Biomedical Research Centre and the European Research Council.
Abstract of Proceedings of the National Academy of Sciences paper
Mass spectrometry imaging (MSI) provides the opportunity to investigate tumor biology from an entirely novel biochemical perspective and could lead to the identification of a new pool of cancer biomarkers. Effective clinical translation of histology-driven MSI in systems oncology requires precise colocalization of morphological and biochemical features as well as advanced methods for data treatment and interrogation. Currently proposed MSI workflows are subject to several limitations, including nonoptimized raw data preprocessing, imprecise image coregistration, and limited pattern recognition capabilities. Here we outline a comprehensive strategy for histology-driven MSI, using desorption electrospray ionization that covers (i) optimized data preprocessing for improved information recovery; (ii) precise image coregistration; and (iii) efficient extraction of tissue-specific molecular ion signatures for enhanced biochemical distinction of different tissue types. The proposed workflow has been used to investigate region-specific lipid signatures in colorectal cancer tissue. Unique lipid patterns were observed using this approach according to tissue type, and a tissue recognition system using multivariate molecular ion patterns allowed highly accurate (>98%) identification of pixels according to morphology (cancer, healthy mucosa, smooth muscle, and microvasculature). This strategy offers unique insights into tumor microenvironmental biochemistry and should facilitate compilation of a large-scale tissue morphology-specific MSI spectral database with which to pursue next-generation, fully automated histological approaches.