First comprehensive atlas of human gene activity released
March 28, 2014
A large international consortium of researchers has produced the first comprehensive, detailed map of the way genes work across the major cells and tissues of the human body. The findings describe the complex networks that govern gene activity, and the new information could play a crucial role in identifying the genes involved with disease.
“Now, for the first time, we are able to pinpoint the regions of the genome that can be active in a disease and in normal activity, whether it’s in a brain cell, the skin, in blood stem cells or in hair follicles,” said Winston Hide, associate professor of bioinformatics and computational biology at Harvard School of Public Health (HSPH) and one of the core authors of the main paper in Nature.
“This is a major advance that will greatly increase our ability to understand the causes of disease across the body.”
The research is outlined in a series of papers published March 27, 2014, two in the journal Nature and 16 in other scholarly journals. The work is the result of years of concerted effort among 250 experts from more than 20 countries as part of FANTOM 5 (Functional Annotation of the Mammalian Genome). The FANTOM project, led by the Japanese institution RIKEN, is aimed at building a complete library of human genes.
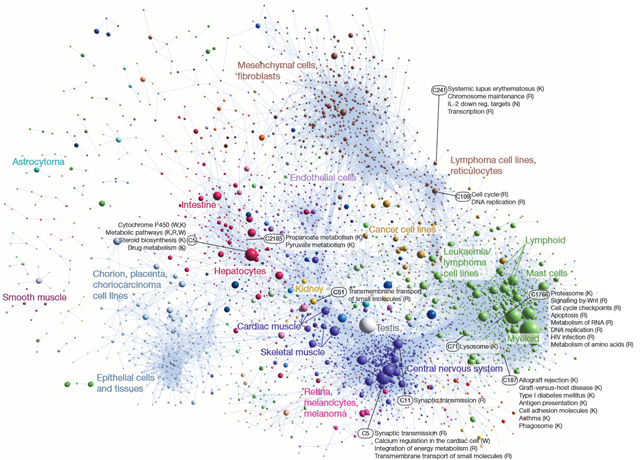
Coexpression clustering of human promoters in FANTOM, showing a portion of a network derived from 4,882 coexpression groups. (Click twice for high-res view.) (Credit: Alistair R. R. Forrest et al./Nature)
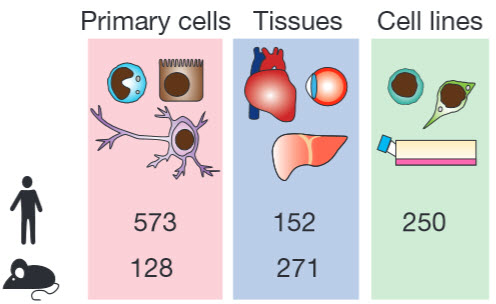
Researchers studied human and mouse cells using a new technology called Cap Analysis of Gene Expression (CAGE), developed at RIKEN, to discover how 95% of all human genes are switched on and off. These “switches” — called “promoters” and “enhancers” — are the regions of DNA that manage gene activity. The researchers mapped the activity of 180,000 promoters and 44,000 enhancers across a wide range of human cell types and tissues and, in most cases, found they were linked with specific cell types.
“We now have the ability to narrow down the genes involved in particular diseases based on the tissue cell or organ in which they work,” said Hide. “This new atlas points us to the exact locations to look for the key genetic variants that might map to a disease.”
Funding for FANTOM 5 came from a research grant from RIKEN and from Innovative Cell Biology.
Abstract of Nature paper
Regulated transcription controls the diversity, developmental pathways and spatial organization of the hundreds of cell types that make up a mammal. Using single-molecule cDNA sequencing, we mapped transcription start sites (TSSs) and their usage in human and mouse primary cells, cell lines and tissues to produce a comprehensive overview of mammalian gene expression across the human body. We find that few genes are truly ‘housekeeping’, whereas many mammalian promoters are composite entities composed of several closely separated TSSs, with independent cell-type-specific expression profiles. TSSs specific to different cell types evolve at different rates, whereas promoters of broadly expressed genes are the most conserved. Promoter-based expression analysis reveals key transcription factors defining cell states and links them to binding-site motifs. The functions of identified novel transcripts can be predicted by coexpression and sample ontology enrichment analyses. The functional annotation of the mammalian genome 5 (FANTOM5) project provides comprehensive expression profiles and functional annotation of mammalian cell-type-specific transcriptomes with wide applications in biomedical research.