Holographic imaging for rapidly sorting stem cells, cancer cells
March 25, 2014
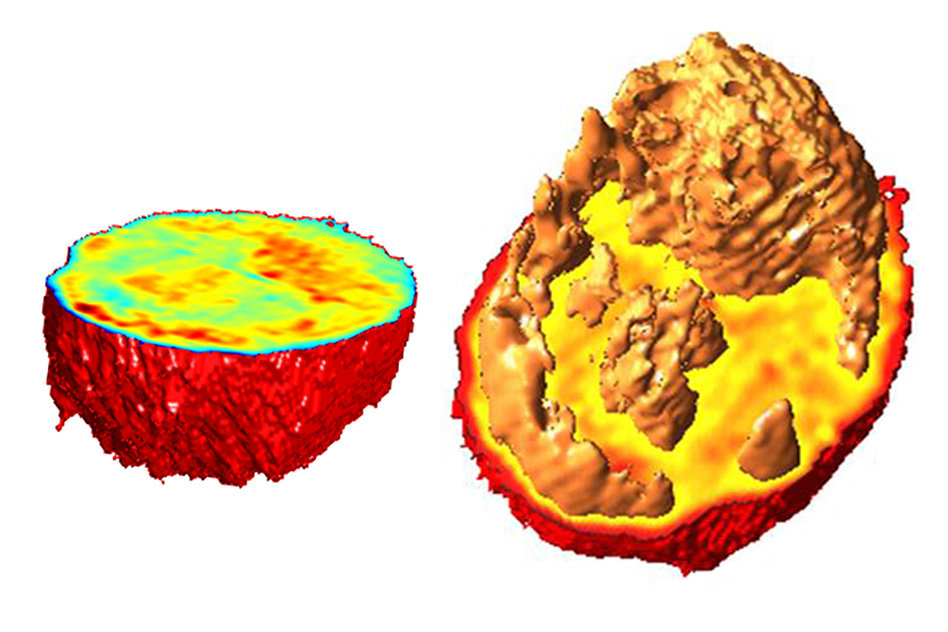
MIT scientists created these 3-D images of living cells based on measurements of their index of refraction. Different colors represent different indices of refraction, which correlate with the cells’ density, allowing for cells to be sorted rapidly in microfluidic channels. (Credit: Niyom Lue)
MIT scientists have developed a way to image cells (without fluorescent markers or other labels) as they flow through a tiny microfluidic channel for sorting.
This is an important step toward cell-sorting systems that could help scientists separate stem cells at varying stages of development, or to distinguish healthy cells from cancerous cells, the scientists say.
Other cell-sorting methods require adding a fluorescent molecule that highlights the cells of interest, but those tags can damage the cells and make them unsuitable for therapeutic uses.
The new method is based on a 2007 microscopy development that allowed the scientists to detail the interior of a living cell in three dimensions, without adding any fluorescent markers or other labels. This technique also revealed key properties, such as the cells’ density.
Sorting stem cells
“Many stem cell applications require sorting of cells at different stages of differentiation. This can be done with fluorescent staining, but once you stain the cells they cannot be used,” says Yongjin Sung, a former postdoc in MIT’s Laser Biomedical Research Center and lead author of a paper describing the technique in the inaugural issue of the journal PRApplied.
“With our approach, you can utilize a vast amount of information about the 3-D distribution of the cells’ mass to sort them.”
Instead of using fluorescent tags, the MIT method analyzes the cells’ index of refraction — a measurement of how much the speed of light is reduced as it passes through a material. Every material has a distinctive index of refraction, and this property can be used, along with cells’ volume, to calculate their mass and density.
Different parts of a cell, including individual organelles, have different indices of refraction, so the information generated by this approach can also be used to identify some of these internal cell structures, such as the nucleus and nucleolus, a structure located within the nucleus.
In the original 2007 version of this technology, known as tomographic phase microscopy, researchers led by the late MIT professor Michael Feld created 3-D images by combining a series of 2-D images taken as laser beams passed through cells from hundreds of different angles. This is the same concept behind CT scanning, which combines X-ray images taken from many different angles to create a 3-D rendering.
Recording continuous flow in real time
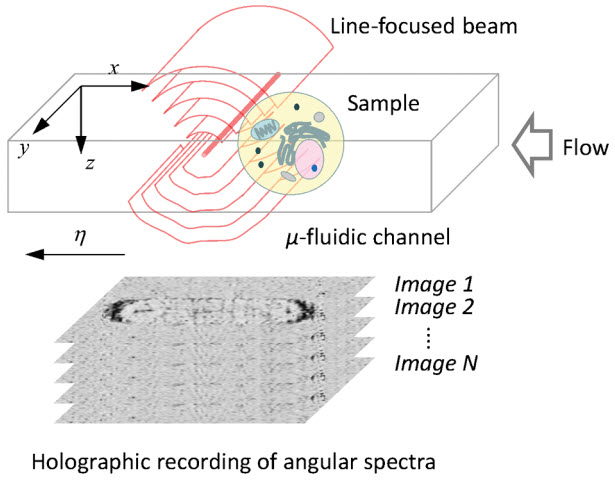
Schematic of three-dimensional holographic imaging of continuously flowing cells (credit: Yongjin Sung et al./PRApplied)
For the new PRApplied study, the MIT researchers collaborated with Daniel Irimia’s lab at Harvard Medical School to adapt the system to image cells as they flow continuously through a microfluidic channel.
Other researchers have tried to do this, but their systems required that the cells be halted at a certain point in the channel to complete the imaging.
“The only moving part in our system is the sample, which will afford great flexibility in miniaturizing the system,” says Sung, who is now an instructor in radiology at Massachusetts General Hospital.
A key feature of the new MIT system is the use of a focused laser beam that can illuminate cells from many different angles, allowing the researchers to analyze the scattered light from the cells as they flow across the beam. Using a technique known as off-axis digital holography, the researchers can instantaneously record both the amplitude and phase of scattered light at each location of the cells.
“As the cell flows across, we can effectively illuminate the entire sample from all angles without having to rotate a light source or the cell,” says former MIT graduate student Niyom Lue, a coauthor of the new paper.
The current system can image about 10 cells per second, but the researchers hope to speed it up to thousands of cells per second, which would make it useful for applications such as sorting stem cells. The researchers also hope to use the system to learn more about how cancer cells grow and respond to different drug treatments.
“This label-free method can look at different states of the cell, whether they are healthy or whether they maybe have cancer or viral or bacterial infections,” says Peter So, an MIT professor of mechanical engineering and biological engineering who is senior author of the new paper. “We can use this technique to look at the pathological state of the cell, or cells under treatment of some drug, and follow the population over a period of time.”
The research was funded by the National Institutes of Health and Hamamatsu Photonics.
Abstract of Physical Review Applied paper
The refractive index of biological specimens is a source of intrinsic contrast that can be explored without any concerns of photobleaching or harmful effects caused by extra contrast agents. In addition, the refractive index contains rich information related to the metabolism of cells at the cellular and subcellular levels. Here, we report a no-moving-parts approach that provides three-dimensional refractive-index maps of biological samples continuously flowing in a microfluidic channel. Specifically, we use line illumination and off-axis digital holography to record the angular spectra of light scattered from flowing samples at high speed. Applying the scalar diffraction theory, we obtain accurate refractive-index maps of the samples from the measured spectra. Using this method, we demonstrate label-free three-dimensional imaging of live RKO human colon cancer cells and RPMI8226 multiple myeloma cells, and obtain the volume, dry mass, and density of these cells from the measured three-dimensional refractive-index maps. Our results show that the reported method, alone or in combination with the existing flow cytometry techniques, shows promise as a quantitative tool for stain-free characterization of a large number of cells.