How to assemble designer biomolecular nanomachines
October 1, 2012

Precise assembly of protein-based nanostructure (credit: Mathias Strackharn et al./Journal of the American Chemical Society)
Assembling novel biomolecular machines requires nanometer precision. Ludwig Maximilian University (LMU) researchers have now found a way to acheive it.
The finely honed tip of the atomic force microscope (AFM) allows one to pick up single biomolecules and deposit them elsewhere with nanometer accuracy. The technique is referred to as Single-Molecule Cut & Paste (SMC&P), and was developed by the research group led by LMU physicist Professor Hermann Gaub.
It was initially only applicable to DNA molecules, but the molecular machines responsible for many of the biochemical processes in cells consist of proteins.
So a practical method of manipulating proteins is required. It would provide novel insights into the workings of living cells and would furnish a way to develop, construct, and utilize designer nanomachines.
Assembling proteins without damage
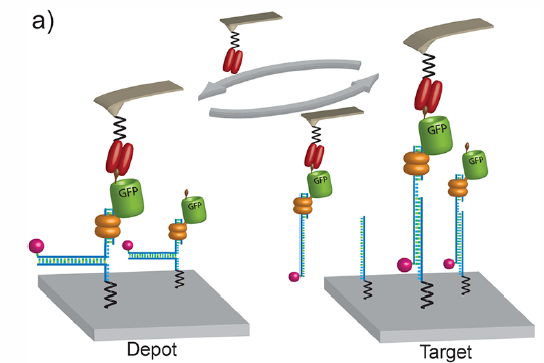
Schematics of the transfer process. The GCN4−GFP− zinc finger construct is bound with high affinity to the anchor DNA and stored via hybridization in the depot area. A single-chain antibody fragment, which is covalently attached to the cantilever tip, seizes the GCN4-tag, and the DNA in unzip geometry opens up when the cantilever is lifted. The GCN4−GFP−zinc finger construct with the anchor DNA can then be transferred to the target site, where the shear geometry bond to the target DNA is stronger than the bond between antibody and GCN4 peptide. The protein construct is deposited in the target area, whereas the antibody on the cantilever is free again and can be reused in the next transfer cycle. (credit: Mathias Strackharn et al./Journal of the American Chemical Society)
In a major step towards this goal, the LMU team has modified the SMC&P method to allow them to take proteins from a storage site and place them at defined locations within a construction area with nanometer precision.
“In liquid medium at room temperature, the “weather conditions” at the nanoscale are comparable to those in a hurricane,” says Mathias Strackharn, first author of the new study. So the molecules being manipulated must be firmly attached to the tip of the AFM and held securely in place in the construction area.
The forces that tether the proteins during transport and assembly must also be weak enough not to cause damage, and must be tightly controlled. To achieve these two goals, the researchers used a combination of antibodies, DNA-binding “zinc-finger” proteins, and DNA anchors.
“We demonstrated the method’s feasibility by bringing hundreds of fluorescent GFP molecules together to form a little green man, like the traffic-light figure that signals to pedestrians to cross the road, but only some micrometers high,” Strackharn explains.
With this technique, functional aspects of complex protein machines — such as how combinations of different enzymes interact, and how close together they must be to perform coupled reactions — can be tested directly. A further goal is to develop artificial multimolecular assemblies modeled on natural “cellulosomes,” which could be used to convert plant biomass into biofuels.
“If we can efficiently build mimics of these ‘enzymatic assembly lines’ by bringing individual proteins together, we could perhaps make a significant contribution to the exploitation of sustainable energy sources.”