Modified atomic force microscope allows for high-speed, high-res imaging of live neurons
March 16, 2015

Extreme closeup of a hippocampal neuron in action (Credit: Max Planck Florida Institute for Neuroscience)
Researchers at the Max Planck Florida Institute for Neuroscience and Kanazawa University (Japan) have imaged structural dynamics of living neurons with unprecedented spatial resolution and speed by using a modified atomic force microscope (AFM).
The AFM is a leading tool for imaging, measuring, and manipulating materials with atomic resolution — on the order of fractions of a nanometer — by scanning (“touching” and “feeling”) its surface with an extremely fine needle (the diameter of the tip is about 5 nanometers) on the surface. However, it can’t be used with neurons and other living cells without damaging the sample, and it’s too slow (it takes many minutes to acquire an image).
Imaging living neurons and other cells
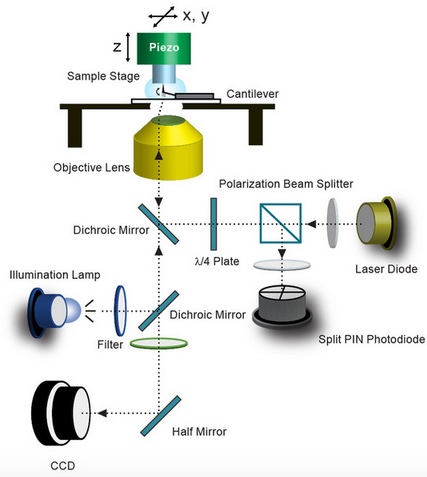
A schematic illustration of the long-tip high-speed atomic force microscope (LT-HS-AFM), including an arc lamp for fluorescence excitation and a CCD camera to capture bright-field and fluorescence images (credit: Mikihiro Shibata/Scientific Reports)
In their March open-access publication in Scientific Reports, Ryohei Yasuda and Mikihiro Shibata at Max Planck Florida Institute for Neuroscience in collaboration with Kanazawa University describe their new long-tip high-speed (LT-HS-AFM) system. It operates at a scanning speed orders of magnitude faster than that of conventional AFM.
To avoid damaging biological samples, using an extremely long, sharp needle attached to a highly flexible plate and a high-speed (800 kHz) light tapping-mode scanning system combined with an extremely soft cantilever to apply low force on the sample. The system uses a wide-area scanner to image large area of mammalian cells (up to ~5 × 5 square micrometers) and optical microscopy to facilitate the positioning of the scanning tip on samples.
The new device images without any sign of cellular damage, it’s optimized for fast scanning (seconds per image) to capture dynamic events in living cells, and has a spatial resolution up to 100 times better than a standard light microscope.
In particular, this study demonstrates the capability to track structural dynamics and remodeling of the cell surface, such as morphogenesis (how the form is created) of filopodia, membrane ruffles, pit formation, or endocytosis in response to environmental stimulants.
Examples of this capability are seen in the movie above, where morphological changes of a finger-like neuronal protrusion in the mature hippocampal neuron are imaged, and in the movie below, where a fibroblast is imaged before and after treatment with insulin hormone, which intensely enhances the ruffling at the leading edge of the cell.
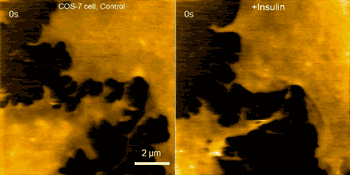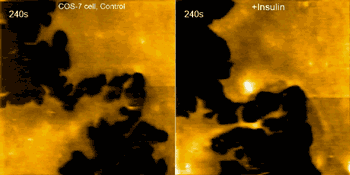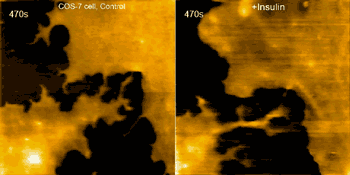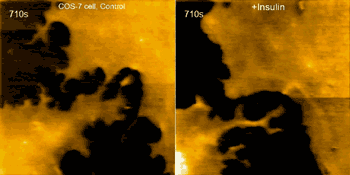
Fibroblast pre/post treatment w/insulin hormone (credit: Max Planck Florida Institute for Neuroscience)
Yasuda says the researchers hope to visualize the morphology of synapses at nanometer resolution and in real time in the near future. Since morphology changes of synapses underlie synaptic plasticity and our learning and memory, this will provide us with many new insights into mechanisms of how neurons store information in their morphology, how it changes synaptic strength, and ultimately how it creates new memory, he suggests.
Abstract of Long-tip high-speed atomic force microscopy for nanometer-scale imaging in live cells
Visualization of morphological dynamics of live cells with nanometer resolution under physiological conditions is highly desired, but challenging. It has been demonstrated that high-speed atomic force microscopy is a powerful technique for visualizing dynamics of biomolecules under physiological conditions. However, application of high-speed atomic force microscopy for imaging larger objects such as live mammalian cells has been complicated because of the collision between the cantilever and samples. Here, we demonstrate that attaching an extremely long (~3 μm) and thin (~5 nm) tip by amorphous carbon to the cantilever allows us to image the surface structure of live cells with the spatiotemporal resolution of nanometers and seconds. We demonstrate that long-tip high-speed atomic force microscopy is capable of imaging morphogenesis of filopodia, membrane ruffles, pit formation, and endocytosis in COS-7, HeLa cells and hippocampal neurons.