New programming language directs DNA to build custom-designed molecules
October 2, 2013
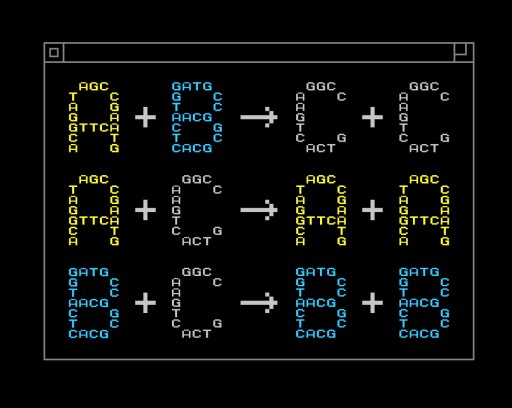
An example of a chemical program. Here, A, B and C are different chemical species. (Credit: Yan Liang/L2XY2.com)
A team led by the University of Washington has developed a programming language to help design chemical-reaction networks (equations that describes how mixtures of chemicals behave).
The objective is to control how DNA molecules build custom-designed molecules in a test tube or cell, which could serve as “smart” drug deliverers or disease detectors at the cellular level, for example.
“We start from an abstract mathematical description of a chemical system, and then use DNA to build the molecules that realize the desired dynamics,” said corresponding author Georg Seelig, a UW assistant professor of electrical engineering and of computer science and engineering.
Currently, when a biologist or chemist makes a certain type of molecular network, the engineering process is complex, cumbersome and hard to repurpose for building other systems.
“I think this is appealing because it allows you to solve more than one problem,” Seelig said. “If you want a computer to do something else, you just reprogram it. This project is very similar in that we can tell chemistry what to do.”
Humans and other organisms already have complex networks of nano-sized molecules that help to regulate cells and keep the body in check. Scientists now are finding ways to design synthetic systems that behave like biological ones with the hope that synthetic molecules could support the body’s natural functions. To that end, a system is needed to create synthetic DNA molecules that vary according to their specific functions.
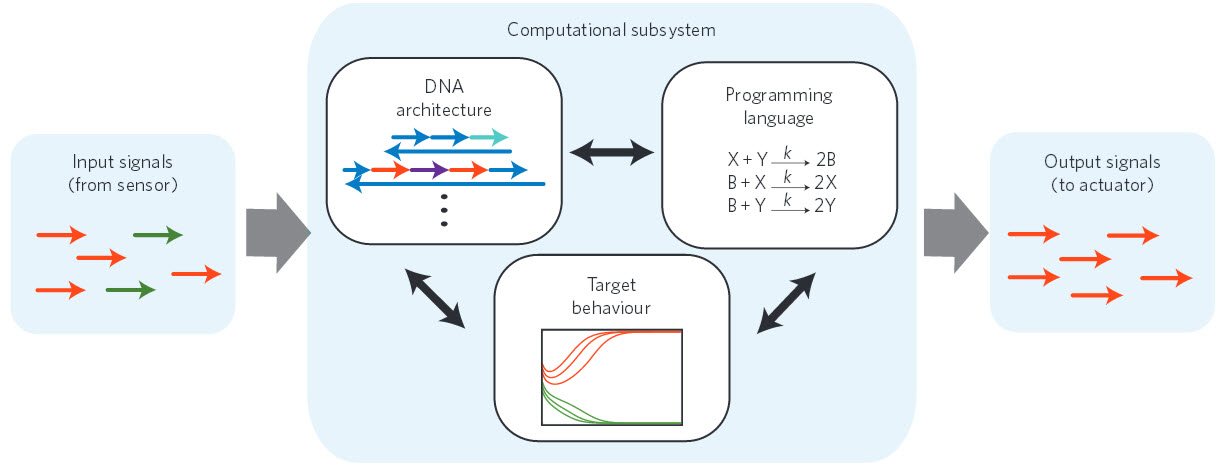
Nucleic acid nanocontroller. A standardized signaling protocol based on short single strands of DNA enables the components of the nanocontroller to communicate with each other. The formalism of chemical-reaction networks serves as a programming language that specifies the desired behavior for the computational subsystem. The target behavior is experimentally realized by the DNA architecture. (Credit: Yuan-Jyue Chen et al., Nature Nanotechnology)
The new approach isn’t ready to be applied in the medical field, but future uses could include using this framework to make molecules that self-assemble within cells and serve as “smart” sensors. These could be embedded in a cell, then programmed to detect abnormalities and respond as needed, perhaps by delivering drugs directly to those cells.
Seelig and colleague Eric Klavins, a UW associate professor of electrical engineering, recently received $2 million from the National Science Foundation as part of a national initiative to boost research in molecular programming. The new language will be used to support that larger initiative, Seelig said. The research was also funded by the Burroughs Wellcome Fund and the National Centers for Systems Biology.
California Institute of Technology; Microsoft Research, and University of California, San Francisco researchers were co-authors of the study.
Abstract of Nature Nanotechnology paper:
Biological organisms use complex molecular networks to navigate their environment and regulate their internal state. The development of synthetic systems with similar capabilities could lead to applications such as smart therapeutics or fabrication methods based on self-organization. To achieve this, molecular control circuits need to be engineered to perform integrated sensing, computation and actuation. Here we report a DNA-based technology for implementing the computational core of such controllers. We use the formalism of chemical reaction networks as a ’programming language’ and our DNA architecture can, in principle, implement any behaviour that can be mathematically expressed as such. Unlike logic circuits, our formulation naturally allows complex signal processing of intrinsically analogue biological and chemical inputs. Controller components can be derived from biologically synthesized (plasmid) DNA, which reduces errors associated with chemically synthesized DNA. We implement several building-block reaction types and then combine them into a network that realizes, at the molecular level, an algorithm used in distributed control systems for achieving consensus between multiple agents.