Programming smart molecules for chemical-based AI
December 19, 2013
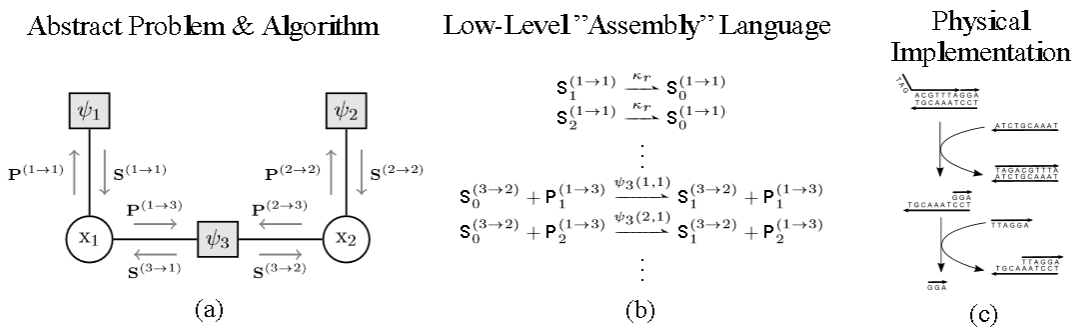
Inference at different levels of abstraction. (a) Factor graph over two random variables. Inference can be performed efficiently by passing messages (shown as gray arrows) between vertices. (b) Message passing implemented at a lower level of abstraction. (c) Schematic representation of DNA strand displacement. (Credit: Nils E. Napp and Ryan Prescott Adams)
Computer scientists at the Harvard School of Engineering and Applied Sciences (SEAS) and the Wyss Institute for Biologically Inspired Engineering at Harvard University have joined forces to put powerful probabilistic reasoning algorithms in the hands of bioengineers.
In a new paper (open access) presented at the recent Neural Information Processing Systems conference, Ryan P. Adams and Nils Napp showed that an important class of artificial intelligence algorithms could be implemented using chemical reactions.
These algorithms, which use a technique called “message passing inference on factor graphs,” are a mathematical coupling of ideas from graph theory and probability. They represent the state of the art in machine learning and are already critical components of everyday tools ranging from search engines and fraud detection to error correction in mobile phones.
‘Smart drugs’
Adams’ and Napp’s work demonstrates that some aspects of artificial intelligence (AI) could be implemented at microscopic scales using molecules. In the long term, the researchers say, such theoretical developments could open the door for “smart drugs” that can automatically detect, diagnose, and treat a variety of diseases using a cocktail of chemicals that can perform AI-type reasoning.
“We understand a lot about building AI systems that can learn and adapt at macroscopic scales; these algorithms live behind the scenes in many of the devices we interact with every day,” says Adams, an assistant professor of computer science at SEAS, whose Intelligent Probabilistic Systems group focuses on machine learning and computational statistics.
“This work shows that it is possible to also build intelligent machines at tiny scales, without needing anything that looks like a regular computer. This kind of chemical-based AI will be necessary for constructing therapies that sense and adapt to their environment. The hope is to eventually have drugs that can specialize themselves to your personal chemistry and can diagnose or treat a range of pathologies.”
Adams and Napp designed a tool that can take probabilistic representations of unknowns in the world (probabilistic graphical models, in the language of machine learning) and compile them into a set of chemical reactions that estimate quantities that cannot be observed directly. The key insight is that the dynamics of chemical reactions map directly onto the two types of computational steps that computer scientists would normally perform in silico to achieve the same end.
Statistical inference by biological reaction pathways and regulatory networks
This insight opens up interesting new questions for computer scientists working on statistical machine learning, such as how to develop novel algorithms and models that are specifically tailored to tackling the uncertainty molecular engineers typically face. In addition to the long-term possibilities for smart therapeutics, it could also open the door for analyzing natural biological reaction pathways and regulatory networks as mechanisms that are performing statistical inference.
Just like robots, biological cells must estimate external environmental states and act on them; designing artificial systems that perform these tasks could give scientists a better understanding of how such problems might be solved on a molecular level inside living systems.
“There is much ongoing research to develop chemical computational devices,” says Napp, a postdoctoral fellow at the Wyss Institute, working on the Bioinspired Robotics platform, and a member of the Self-organizing Systems Research group at SEAS. Both groups are led by Radhika Nagpal, the Fred Kavli Professor of Computer Science at SEAS and a Wyss core faculty member. At the Wyss Institute, a portion of Napp’s research involves developing new types of robotic devices that move and adapt like living creatures.
“What makes this project different is that, instead of aiming for general computation, we focused on efficiently translating particular algorithms that have been successful at solving difficult problems in areas like robotics into molecular descriptions,” Napp explains. “For example, these algorithms allow today’s robots to make complex decisions and reliably use noisy sensors. It is really exciting to think about what these tools might be able to do for building better molecular machines.”
Indeed, the field of machine learning is revolutionizing many areas of science and engineering. The ability to extract useful insights from vast amounts of weak and incomplete information is not only fueling the current interest in “big data,” but has also enabled rapid progress in more traditional disciplines such as computer vision, estimation, and robotics, where data are available but difficult to interpret. Bioengineers often face similar challenges, as many molecular pathways are still poorly characterized and available data are corrupted by random noise.
Using machine learning, these challenges can now be overcome by modeling the dependencies between random variables and using them to extract and accumulate the small amounts of information each random event provides.
“Probabilistic graphical models are particularly efficient tools for computing estimates of unobserved phenomena,” says Adams. “It’s very exciting to find that these tools map so well to the world of cell biology.”
Abstract of Neural Information Processing Systems paper
Recent work on molecular programming has explored new possibilities for computational abstractions with biomolecules, including logic gates, neural networks, and linear systems. In the future such abstractions might enable nanoscale devices that can sense and control the world at a molecular scale. Just as in macroscale robotics, it is critical that such devices can learn about their environment and reason under uncertainty. At this small scale, systems are typically modeled as chemical reaction networks. In this work, we develop a procedure that can take arbitrary probabilistic graphical models, represented as factor graphs over discrete random variables, and compile them into chemical reaction networks that implement inference. In particular, we show that marginalization based on sum-product message passing can be implemented in terms of reactions between chemical species whose concentrations represent probabilities. We show algebraically that the steady state concentration of these species correspond to the marginal distributions of the random variables in the graph and validate the results in simulations. As with standard sum-product inference, this procedure yields exact results for tree-structured graphs, and approximate solutions for loopy graphs.