Sequencing the Connectome
October 25, 2012
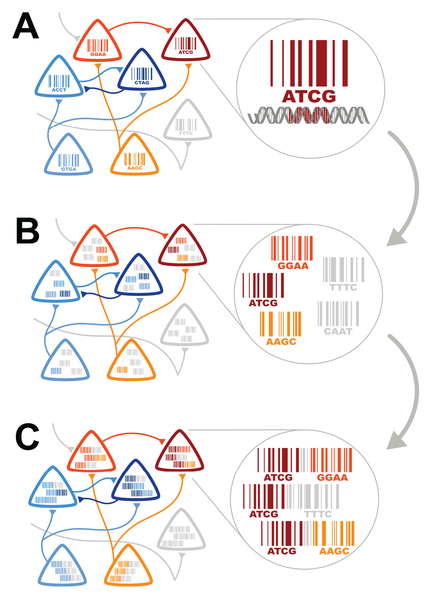
Converting connectivity into a sequencing problem can be broken down conceptually into three components. (A) Label each neuron with a unique sequence of nucleotides — a DNA “barcode.” (B) Associate barcodes from synaptically connected neurons with one another, so that each neuron can be thought of as a “bag of barcodes” — copies of its own “host” barcode and copies of “invader” barcodes from synaptic partners. (C) Join host and invader barcodes into barcode pairs. These pairs can be subjected to high-throughput sequencing. (Credit: Anthony M. Zador et al./PLoS Biology)
A team of neuroscientists led by Professor Anthony Zador, Ph.D., of Cold Spring Harbor Laboratory have proposed a revolutionary new way to create a connectivity map (“connectome”) of the whole brain of the mouse at the resolution of single neurons: high-throughput DNA sequencing.
The only current method for obtaining the connectome with high precision relies on laboriously examining individual cell-to-cell contacts (synapses) in electron microscopes, which is slow, expensive and labor-intensive. (See A circuit diagram of the mouse brain.)
This reconstruction of serial electron micrographs has yielded what to date is the only complete connectome, that of C. elegans (a nemotode or roundworm). However, even for this simple nervous system, the reconstruction required a heroic effort — more than 50 person-years of labor to collect and analyze the images.
The appeal of using sequencing is that its scale and speed — sequencing billions of nucleotides per day is now routine — is a natural match to the complexity of neural circuits. And it’s getting faster exponentially.
An inexpensive high-throughput technique for establishing circuit connectivity at single neuron resolution could transform neuroscience research, the open-access PLoS Biology paper says.
Barcoding synapses
“Our method renders the connectivity problem in a format in which the data are readable by currently available high-throughput genome sequencing machines,” says Zador. “We propose to do this via a process we’re now developing, called BOINC: barcoding of individual neuronal connections.”
The proposal comes at a time when a number of scientific teams in the U.S. are progressing in their efforts to map connections in the mammalian brain. These efforts use injections of tracer dyes or viruses to map neuronal connectivity at a “mesoscopic” scale — a mid-range resolution that makes it possible to follow neural fibers between brain regions. Other groups are scaling up approaches based on electron microscopy.
Zador’s team wants to trace connectivity “beyond the mesoscopic,” at the level of synaptic contacts between pairs of individual neurons, throughout the brain. The BOINC barcoding technique, now undergoing proof-of-concept testing, will be able, says Zador, “to provide immediate insight into the computations that a circuit performs.”
In practice, he adds, most neural computations are not currently understood at this level of precision, partly because detailed circuit information is not available for mammals. The BOINC method promises to be much faster and cheaper than approaches based on electron microscopy, Zador says. (The paper includes a cost analysis, with a possible $1,000 mouse cortical connectome,)
How BOINC would work
- Label each neuron with a specific DNA barcode, consisting of just 20 random DNA “letters.” (A barcode consisting of even 20 random nucleotides can uniquely label 420 = 1012 neurons — 10,000 times more than the number of neurons [<108] in a mouse brain.)
- Look at neurons that are synaptically connected and associate their respective barcodes with one other. One way to do this is by using a virus (such as the pseudorabies virus) that can move genetic material across synapses. “To share barcodes across synapses, the virus must be engineered to carry the barcode within its own genetic sequence,” explains Zador. “After the virus spreads across synapses, each neuron effectively ends up as a bag of barcodes, comprising its own code and those from synaptically coupled partners.”
- Join barcodes from synaptically connected neurons to make single pieces of DNA, which can then be read via existing high-throughput DNA sequencing methods. These double-barcode sequences can then be analyzed computationally to reveal the synaptic wiring diagram of the brain. (Barcodes are joined in vivo, so there is no need to isolate individual neurons prior to extracting DNA.)
If BOINC succeeds in its current proof-of-concept tests, it will offer a dramatically inexpensive and rapid means of assembling a connectome, even of the complex brains of mammals, says Zador.
This work was funded by grants from the NIH and the Paul Allen Family Foundation.