Web apps for bioinformatics
July 24, 2012
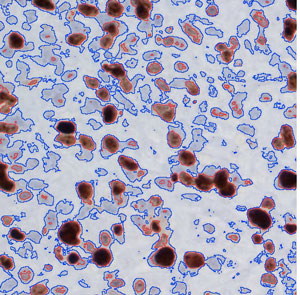
This snapshot is the result of using an Imagejs module to determine how fast brain cancer cells are growing. The darks blots are the nuclei of cells dividing as part of the high-speed abnormal growth seen in tumors. (Credit: UAB)
A University of Alabama at Birmingham (UAB) team has developed ImageJS, a free app system that analyzes tissue images.
The ImageJS module enables pathologists to drag a digitized pathology slide into a Web app and analyze it for malignancy based on color. Cancer cells change color when exposed to standard dyes. It is available from the Google Chrome App store, Google Code and Github.
ImageJS is the first in a series coming from Jonas Almeida, Ph.D., director of the Division of Informatics in the UAB School of Medicine Department of Pathology.
Future modules will seek to perform genomics analysis, make the system capable of cloud computing, and enable doctors to compare their patient’s data to similar cases stored in national databases. Such comparisons promise to increase diagnostic accuracy and rule out treatments that clash with a person’s genetic signature.
Public resources such as The Cancer Atlas, Gene Expression Omnibus and 1,000 Genomes Project have been generating massive data sets for years, but researchers are just beginning to harness them with Web apps that improve health-care delivery. To tap this potential, experts need apps that are neither overly complex nor require unsafe downloads of desktop applications.
The promise of ImageJS, Almeida says, depends on whether or not pathologists partner with biostatisticians at other institutions to write modules that add value or resolve their specific problems. If that happens, ImageJS could come to resemble the kind of social coding communities that surround smart phones, in this case to improve care and empower research.
Replacing complex Java code with JavaScript
The “ImageJS” name pays homage to the “ImageJ”, an image-analysis program pioneered by the National Institutes of Health. ImageJ was written in the Java language and required considerable programming efforts and security-breaching software to make the small changes needed to work with a hospital’s system. Because hospital systems often prevent just such software downloads, its use has been limited there.
Java also posed challenges for the mobile Web and tablet platforms, which seldom support code written in languages other than the browser’s native JavaScript, says Almeida. In contrast, the new ImageJS, a collection of Web-based algorithms written in JavaScript, enables researchers to change apps to fit their needs with a few lines of code and repeat each other’s informatics experiments.
Most important, ImageJS code migrates from the code repository to the browser and eliminates the risks of travel-related data damage and exposure that violates patient privacy.
Web 3.0 data compatibility
The central barrier to Web 3.0, an approaching era when all computers act as one to achieve new levels of collaboration and computing power, is that data compiled to date worldwide has been locked in silos that can’t be searched, shared or analyzed.
While researchers hope to translate existing databases into the universal Web 3.0 data formats that will allow such sharing, they are also starting fresh. Almeida’s team, for instance, will be attempting over the next year to build a database that offers future ImageJS users the option to store images and related analysis in a resource description framework or RDF.
Once data is stored in this format, it can be found by queries written in SPARQL, a simple computer language that can be learned by most people in a couple of hours. That is extremely important, says Almeida, because the next generation of bioinformatics must not remain the province of programmers.
Instead, the researchers running the clinical trials and translational studies must write their own programs, which would add “stunning, new analytic value” to their research and feed into the new global databases.
The UAB team’s work proceeds alongside other academic efforts, such as Tetherless World Constellation at Rensselaer Polytechnic Institute and ventures such as LinkedLifeData, Sindice and Hadoop.