Zooming in on bacterial weapons in 3D
May 23, 2012
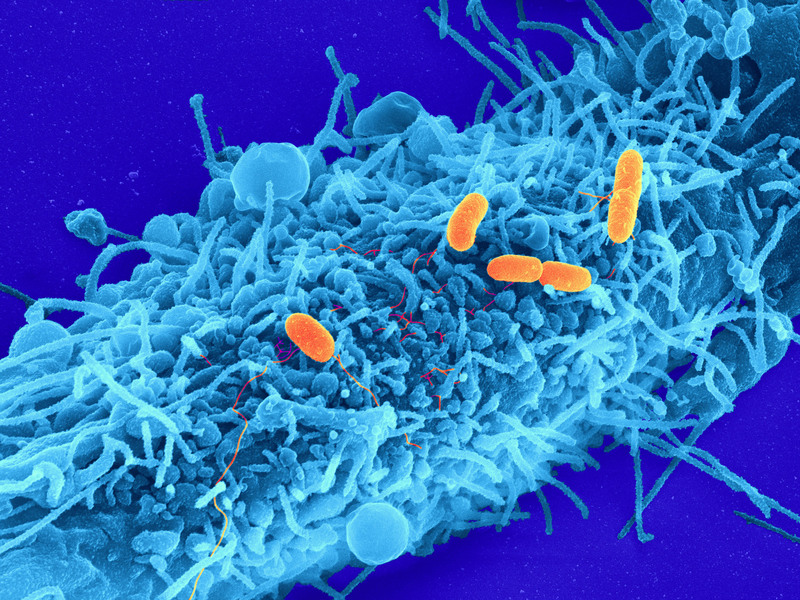
Bacterial infection of host cells: Pathogens of the type Salmonella typhimurium (orange) establish contact to a human host cell (blue) (credit: Christian Goosmann, Diane Schad, Rashmi Gupta and Michael Kolbe/Max Planck Institute)
Imaging of the structure of bacteria’s injection needles at atomic resolution has been achieved by researchers at the Max Planck Institute for Biophysical Chemistry in cooperation with colleagues at the Max Planck Institute for Infection Biology and the University of Washington.
Their findings might contribute to drug tailoring and the development of strategies which specifically prevent the infection process.
Hundreds of tiny hollow needles sticking out of the bacterial membrane — it is a treacherous tool that makes pathogens causing plague or cholera so dangerous.
Together with a base, embedded in the membrane, these miniature syringes constitute the “type III” secretion system — an injection apparatus through which the pathogens introduce molecular agents into their host cell. There, these substances manipulate essential metabolic processes and disable the immune defence of the infected cells.
The consequences are fatal, as the pathogens can now spread within the organism without hindrance. To date, traditional antibiotics are prescribed to fight the infection. However, as some bacterial strains succeed in developing resistances, researchers worldwide seek to discover more specific drugs.
The exact structure of the 60 to 80 nanometer (60 to 80 millionths of a millimeter) long and about eight nanometet wide needles has so far been unknown. Classical methods such as X-ray crystallography or electron microscopy failed or yielded wrong model structures. Not crystallisable and insoluble, the needle resisted all attempts to decode its atomic structure.
Adam Lange and Stefan Becker at the Max Planck Institute for Biophysical Chemistry together with a team of physicists, biologists and chemists chose a completely novel approach. In cooperation with David Baker at the University of Washington, and Michael Kolbe at the Max Planck Institute for Infection Biology, the scientists successfully combined the production of the needle in the laboratory with solid-state NMR spectroscopy, electron microscopy, and computer modelling.
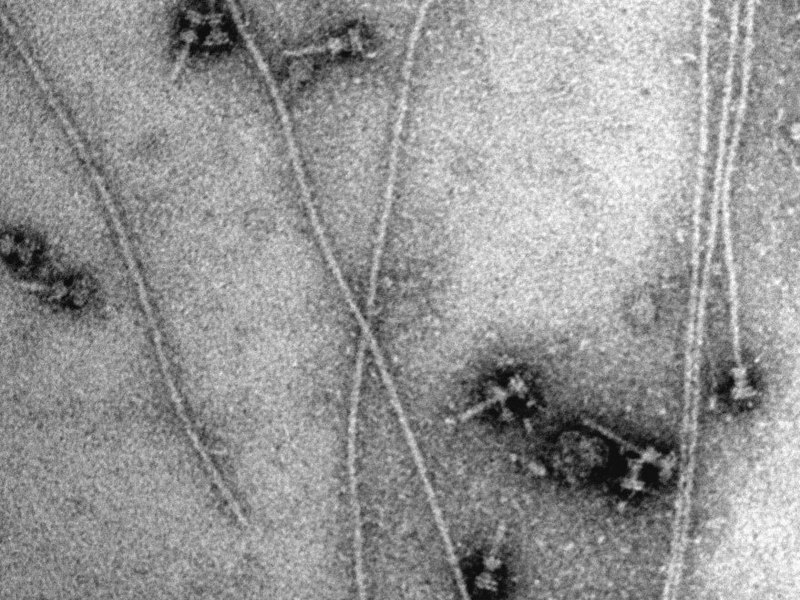
Syringes isolated from Shigella flexneri. Adding soluble needle protein leads to a spontaneous elongation of some needles. The bar corresponds to 100 nanometers (1 nanometer corresponds to a millionth millimeter). (Credit: Christian Goosmann, Michael Kolbe/Mac Planck Institute)
The researchers deciphered the structure of the needle atom by atom and visualised its molecular architecture for the first time in the angstrom range (one tenth of a nanometer).
This required progresses in several fields. “We have made big steps forward concerning sample production as well as solid-state NMR spectroscopy,” says Adam Lange. “Finally, we were also able to use one of the presently most powerful solid-state NMR spectrometers in Christian Griesinger’s NMR-based Structural Biology Department at our Institute.” With 20 tesla, the magnetic field of this 850 megahertz spectrometer is about 400,000 times as strong as that of the Earth.
“We were surprised to see how the needles are constructed,” says Lange. As expected, the needles of pathogens causing diseases as diverse as food poisoning, bacterial dysentery, or the plague show striking similarities. However, in contrast to prevailing assumptions, the similarities are found in the inner part of the needles whereas the surface is astonishingly variable. According to the scientist, this variability might be a strategy of the bacteria to evade immune recognition by the host. Changes on the surface of the needle make it difficult for the host’s immune system to recognize the pathogen.
The discovery of their structure in atomic detail not only enables researchers to gain new insights into how these pathogens outwit their host cells, it also offers the prospect to block the syringe assembly and the delivery of the bacterial factors using tailored molecules.
Such substances, referred to as antiinfectives, could act more specifically and much earlier during infection than traditional antibiotics. “Thanks to our new technique, we can produce large amounts of needles in the lab. Our aim is now to develop a high-throughput method. This will allow us to search for new agents that prevent the formation of the needle,” explains Stefan Becker.
Ref.: Antoine Loquet et al., Atomic Model of the Type III Secretion System Needle, Nature, 2012, DOI: 10.1038/nature11079