A new ‘off’ switch for neurons
April 28, 2014
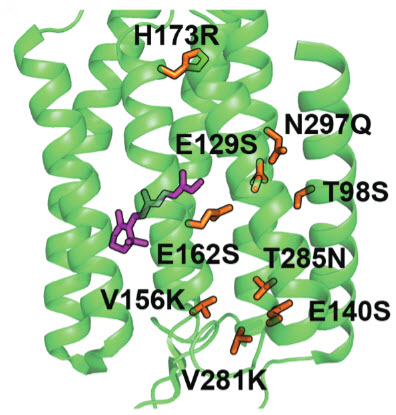
Modified light-activated channelrhodopsin protein structure. The nine mutations to achieve neuron silencing are shown in orange. (Credit: Andre Berndt et al./Science)
Howard Hughes Medical Institute (HHMI) scientists have found a way to silence (turn off) individual neurons, using optogenetics, which previously could only turn on neurons.
Optogenetics is a method used by neuroscientists to study how specific neurons work. They shine LED or laser light on genetically modified neurons (light-activated) proteins (such as channelrhodopsin) that function as ion channels, which then turn on the neurons in which these ion channels are expressed.
HHMI scientist Karl Deisseroth and his colleagues at Stanford University published their new findings April 25, 2014 in the journal Science. “We’re excited about this increased light sensitivity of inhibition in part because we think it will greatly enhance work in large-brained organisms like rats and primates,” he says.
Technical details
First discovered in unicellular green algae in 2002, channelrhodopsins function as photoreceptors that guide the microorganisms’ movements in response to light. In a landmark 2005 study, Deisseroth and his colleagues described a method for expressing the light-sensitive proteins in mouse neurons. By shining a pulse of blue light on those neurons, the researchers showed they could reliably induce the ion channel at channelrhodopsin’s core to open up, allowing positively charged ions to rush into the cell and trigger action potentials. Channelrhodopsins have since been used in hundreds of research projects investigating the neurobiology of everything from cell dynamics to cognitive functions.
A few years later came the deployment of halorhodopsins, light-sensitive proteins selective for the negatively charged ion chloride. These proteins, derived from halobacteria, provided researchers with a tool for the light-controlled inactivation of neurons. A major limitation of these proteins, however, is their inefficiency. Unlike channelrhodopsin, halorhodopsin is an ion pump, meaning that only one chloride ion moves across the neuron’s membrane per photon of light. “What that translates into is you get partial inhibition,” Deisseroth says. “You can inhibit neurons, but in the living animal it’s not always complete.”
Searches for a naturally occurring light-sensitive channel with a pore permeable to negatively charged ions have come up empty handed. “We searched,” Deisseroth says. “We did big genomic searches and found many interesting channelrhodopsins and lots of pumps, but we never found an inhibitory channel in nature.”
The team’s fruitless exploration led them to try modifying the molecular structure of channelrhodopsin so that its pore would shuttle negative ions into the cell. “To do that you need to know what the channel pore looks like at the angstrom level,” Deisseroth says. “What we really needed was the high-resolution crystal structure.” In 2012, working with a group in Japan, Deisseroth and his colleagues captured the structure of a chimera of channelrhodopsin called C1C2 using X-ray crystallography.
A molecular analysis of channelrhodopsin’s pore suggested that swapping out certain negatively charged amino acid residues lining the pore with positive residues could reverse the electrostatic potential of the channel, making it more conductive to negatively charged ions such as chloride.
To achieve this molecular switcheroo, the researchers performed dozens of single site-directed mutations. Several mutations conferred selectivity for chloride, but the channels failed to conduct current. So, the team screened hundreds of combinations of mutations. “In a systematic process we found first a combination of four mutations, and then a group of five mutations, that seemed to change selectivity,” says Deisseroth. “We put those together into a nine-fold mutated channel and that one, amazingly, was chloride selective.”
Not only does the new channel — dubbed iC1C2 for “inhibitory C1C2” — allow the selective passage of chloride ions, it greatly reduces the likelihood of action potentials by making the neuron more “leaky,” a function not possible in ion pumps like halorhodopsin.
Deisseroth’s team made a final mutation to a cysteine residue in iC1C2 that makes the channel both bi-stable and orders of magnitude more sensitive to light. When activated by blue light, the mutated channels remain open for up to minutes at a time, while exposing the channels to red light makes them close quickly. This level of long-term control is useful in developmental studies where events play out over minutes to hours.
The long times for channel opening also mean that neurons can essentially integrate chloride currents over longer time scales and, therefore, weaker light can be used to inhibit the neurons. Increased light sensitivity translates to less light-induced damage to neural tissue, the ability to reach deep brain structures, and the possibility of controlling brain functions that involve large regions of the brain.
“This is something we’ve sought for many years and it’s really the culmination of many streams of work in the lab — crystal structure work, mutational work, behavioral work —all of which have come together here,” Deisseroth says.
Abstract of Science paper
Using light to silence electrical activity in targeted cells is a major goal of optogenetics. Available optogenetic proteins that directly move ions to achieve silencing are inefficient, pumping only a single ion per photon across the cell membrane rather than allowing many ions per photon to flow through a channel pore. Building on high-resolution crystal-structure analysis, pore vestibule modeling, and structure-guided protein engineering, we designed and characterized a class of channelrhodopsins (originally cation-conducting) converted into chloride-conducting anion channels. These tools enable fast optical inhibition of action potentials and can be engineered to display step-function kinetics for stable inhibition, outlasting light pulses and for orders-of-magnitude-greater light sensitivity of inhibited cells. The resulting family of proteins defines an approach to more physiological, efficient, and sensitive optogenetic inhibition.