A super-resolution window into the center of a cell
October 11, 2013
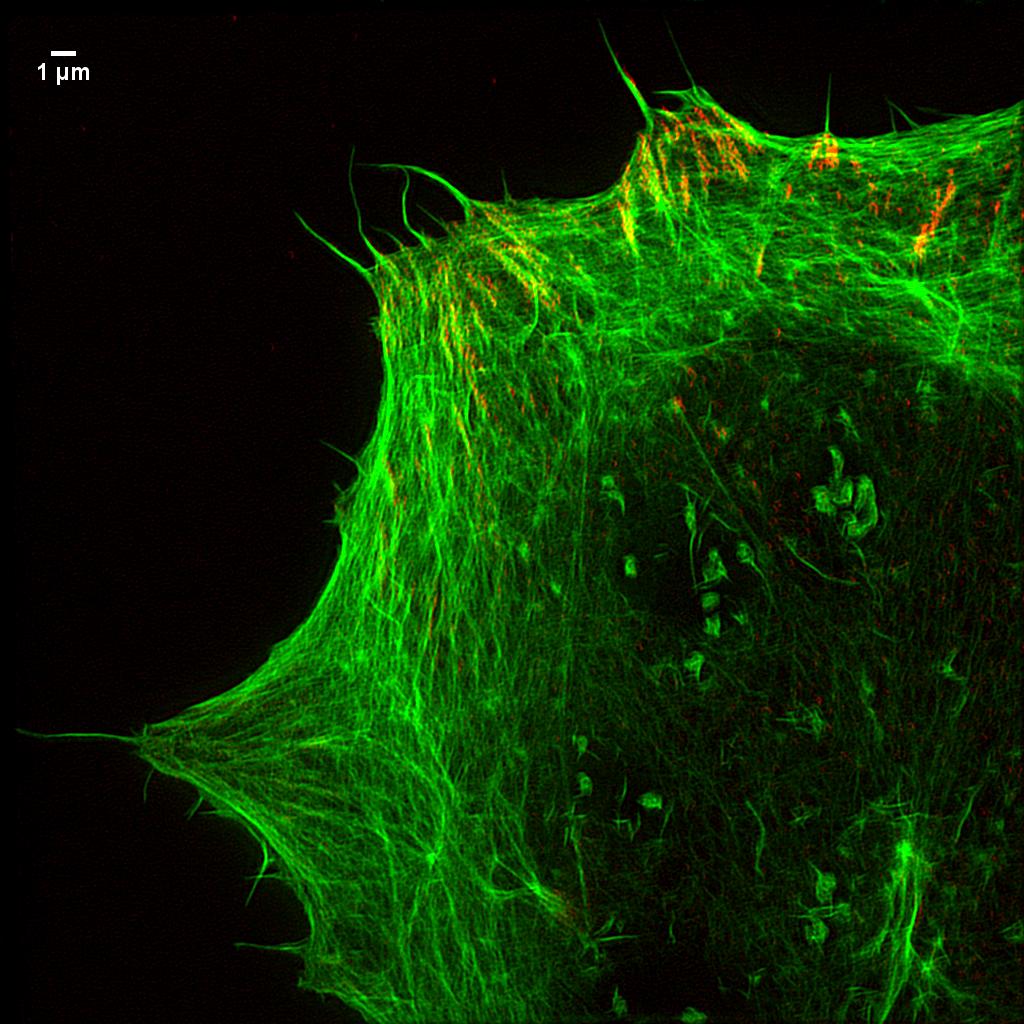
Focal Adhesions and actin. This is a super-resolution image, collected by structured illumination super-resolution microscopy of the actin cytoskeleon and small adhesions, termed focal adhesions. The cell uses focal adhesions to attach to its surroundings. (Credit: Queen Mary University of London)
A new microscopic technique that can see tiny structures inside the “control center” of the cell for the first time has been developed by researchers at Queen Mary University of London,
It represents a major advance for cell biologists because it will allow them to investigate structures deep inside the cell, such as viruses, bacteria, and parts of the nucleus in depth.
Recent advances in optical physics have made it possible to use fluorescent microscopy to study complex structures smaller than the Abbe limit of 200 nanometers (nm) — around 500 times smaller than the width of a human hair.
These methodologies are called super-resolution microscopy.
The drawback of such techniques is that they can only produce very clear images of structures that are at the bottom of the cell. Since the nucleus — the cell’s ‘control center’ — is in the middle of the cell and bacterial and viral infections can happen anywhere in the cell, this technique has considerable limitations for biologists.
The newly developed imaging system, making it possible to image structures as small as 80nm or less anywhere in the cell. It was developed by Dr Neveen Hosny, a bioengineer working with Dr. Ann Wheeler, Head of Imaging at Queen Mary’s Blizard Institute.and Professor Martin Knight in the School of Engineering and Materials Science.
Super-resolution microscopy
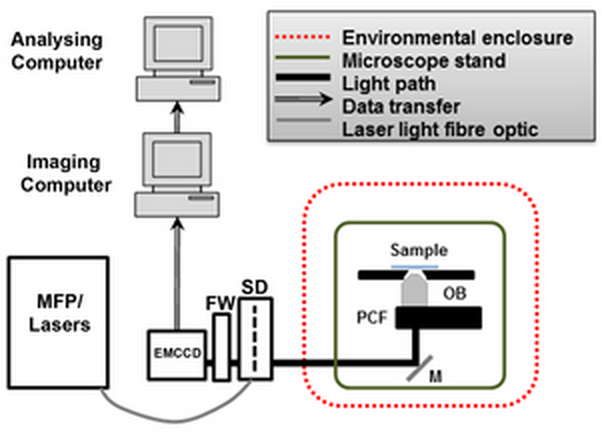
Diagram showing the configuration of the SDSI microscope, abbreviations are as follows. EMCCD = Electron multiplied Charge Coupled Device camera. FW = Filter wheel. SD = Yokagawa CSUX1 spinning disk, M = mirror. PCF = Piezo coupled focus feedback unit. OB = Objective. (Credit: Queen Mary University of London)
The Spinning Disk Statistical Imaging (SDSI) system microscope produces focused images at high speed because it has a disk with an array of tiny holes in it that remove the out-of-focus light, according to Wheeler.
“We have combined this microscope with new fluorescent probes, which switch between a bright and dark state rapidly, she said. “This system is now allowing us to see structures three times smaller than could usually be seen using standard light microscopes.
“We have been able to visualize chromatin, which is the protein structure that controls DNA expression and the nuclear envelope. We have also used the method to get images of focal adhesions — sub-cellular macromolecules that the cell uses to attach to its environment.
“Although it was previously possible to see these structures, our method provides a greater degree of detail.
“It also allows us to look at protein complexes that are smaller than 200nm in the nucleus, which hasn’t been done before.”
“Super resolution microscopy is a major step forward and we are looking forward to using this technology in a wide range of applications from stem cell behavior to understanding arthritis or the development of nanomedicine,” said Knight.
Wheeler has worked with colleagues across Queen Mary to make the technique cost-effective and easy to use for scientists who are not experts in optical physics.
“We will be continuing to develop the technology to improve the fluorescent probes used for this technique and also applying it to cellular processes such as invasion in cancer,” Wheeler added.