Steering the epigenome to turn specific genes on
April 16, 2015
(credit: Human Epigenome Project)
Duke University researchers have developed a new method to precisely control when genes are turned on and active: by manipulating the epigenome — the web of proteins that supports and controls gene activity and a current hot topic in cancer research.
The researchers say having the ability to steer the epigenome will help them explore the roles that particular promoters and enhancers play in cell fate or the risk for genetic disease, and it could provide a new avenue for gene therapies and guiding stem cell differentiation.
The study appeared online April 6 in Nature Biotechnology.
Master control
“The epigenome is everything associated with the genome other than the actual genetic sequence, and is just as important as our DNA in determining cell function in healthy and diseased conditions,” said Charles Gersbach, assistant professor of biomedical engineering at Duke.
“That becomes immediately obvious when you consider that we have over 200 cell types, and yet the DNA in each is virtually the same. The epigenome determines which genes each cell activates and to what degree.”
This genetic puppetmaster consists of DNA-packaging proteins called histones and a host of chemical modifications — either to these histones or the DNA itself — that help determine whether a gene is on or off.
But Gersbach’s team didn’t have to modify the genes themselves to gain some control.
Promoters and enhancers
“Next to every gene is a DNA sequence called a promoter that controls its activity,” explained Gersbach. “But there’s also many other pieces of the genome called enhancers that aren’t next to any genes at all, and yet they play a critical role in influencing gene activity, too.”
Timothy Reddy, assistant professor of biostatistics and bioinformatics at Duke, has spent the better part of a decade mapping millions of these enhancers across the human genome. There has not, however, been a good way to find out exactly what each one does. An enhancer might affect a gene next door or several genes across the genome — or maybe none at all.
To activate these enhancers and see what they do, Reddy thought perhaps he could chemically alter the histones at the enhancers to turn them on.
“There are already drugs that will affect enhancers across the whole genome, but that’s like scorching the earth,” said Reddy. “I wanted to develop tools to go in and modify very specific epigenetic marks in very specific places to find out what individual enhancers are doing.”
Hijacking CRISPR, the gene cut-and-paste system
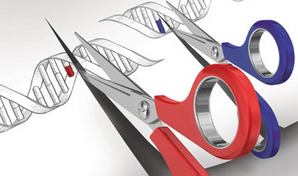
Genome engineering with CRISPR/Cas nucleases. Different Cas9 enzymes (different colored scissors) target different locations in the genome. (credit: Feng Zhang/MIT)
Reddy found that specificity by teaming up with Gersbach, his neighbor within Duke’s Center for Genomic and Computational Biology, who specializes in a gene-targeting system called CRISPR.
Originally discovered as a natural antiviral system in bacteria, researchers have hijacked the system over the past few years and are now using it to cut and paste DNA sequences in the human genome.
But for this epigenome editing application, Gersbach silenced the DNA-cutting mechanism of CRISPR and used CRISPR solely as a targeting system to deliver an enzyme (acetyltransferase) to specific promoters and enhancers and alter the DNA’s packaging at specific sites.
Gersbach and Reddy put their artificial epigenetic agent to the test by targeting a few well-studied gene promoters and enhancers. While these histone modifications have long been associated with gene activity, it wasn’t clear if they were enough to turn genes on. And though Gersbach and Reddy had previously used other technologies to activate gene promoters, they had not successfully activated enhancers.
To the duo’s great surprise, not only did the agent activate the gene promoters, it turned on the adjacent genes better than their previous methods. Equally surprising was that it worked on enhancers as well: they could turn on a gene — or even families of genes — by targeting enhancers at distant locations in the genome, something that their previous gene activators could not do.
Probing enhancers to learn how they relate to disease or drug-therapy response
But the real excitement from their results is an emerging ability to probe millions of potential enhancers in a way never before possible.
“Some genetic diseases are straightforward — if you have a mutation within a particular gene, then you have the disease,” said Isaac Hilton, postdoctoral fellow in the Gersbach Lab and first author of the study.
“But many diseases, like cancer, cardiovascular disease, or neurodegenerative conditions, have a much more complex genetic component. Many different variations in the genome sequence can affect your risk of disease, and this genetic variation can occur in these enhancers that Tim has identified, where they can change the levels of gene expression. With this technology, we can explore what exactly it is that they’re doing and how it relates to disease or response to drug therapies.”
Gersbach added, “Not only can you start to answer those questions, but you might be able to use this technique for gene therapy to activate genes that have been abnormally silenced or to control the paths that stem cells take toward becoming different types of cells. These are all directions we will be pursuing in the future.”
This work was supported by the National Institutes of Health and the National Science Foundation.
Abstract of Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers
Technologies that enable targeted manipulation of epigenetic marks could be used to precisely control cell phenotype or interrogate the relationship between the epigenome and transcriptional control. Here we describe a programmable, CRISPR-Cas9-based acetyltransferase consisting of the nuclease-null dCas9 protein fused to the catalytic core of the human acetyltransferase p300. The fusion protein catalyzes acetylation of histone H3 lysine 27 at its target sites, leading to robust transcriptional activation of target genes from promoters and both proximal and distal enhancers. Gene activation by the targeted acetyltransferase was highly specific across the genome. In contrast to previous dCas9-based activators, the acetyltransferase activates genes from enhancer regions and with an individual guide RNA. We also show that the core p300 domain can be fused to other programmable DNA-binding proteins. These results support targeted acetylation as a causal mechanism of transactivation and provide a robust tool for manipulating gene regulation.