Atomically precise self-assembling nanoscale fibers mimic living cells
August 5, 2014
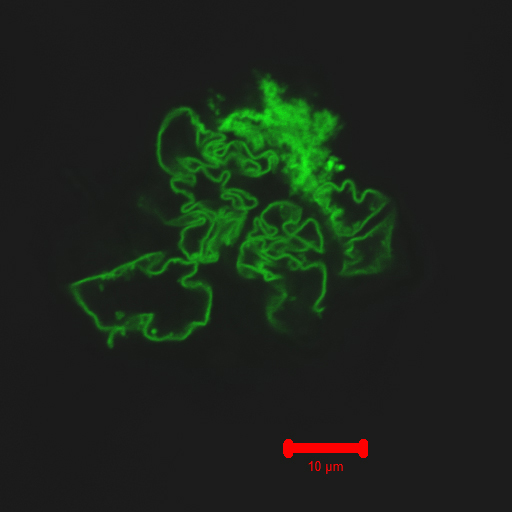
Carnegie Mellon chemists have created a self-assembling nano fiber using green fluorescent protein (credit: Carnegie Mellon University)
Carnegie Mellon University researchers have developed a novel method for creating self-assembled protein/polymer nanostructures that resemble fibers found in living cells.
The work offers a promising new way to fabricate materials for drug delivery and tissue engineering applications.
The findings were published in the July 28 issue of Angewandte Chemie International Edition.
“We have demonstrated that, by adding flexible linkers to protein molecules, we can form completely new types of aggregates,” said Tomasz Kowalewski, professor of chemistry in Carnegie Mellon’s Mellon College of Science.
“These aggregates can act as a structural material to which you can attach different payloads, such as drugs. In nature, this protein isn’t close to being a structural material.”
“The GFP molecules appeared to self-assemble….”
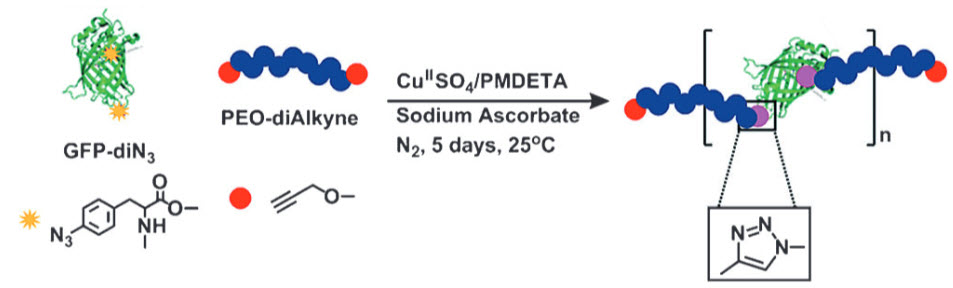
Self-assembling fiber synthesis (credit: Carnegie Mellon University)
The building blocks of the fibers are a few modified green fluorescent protein (GFP) molecules linked together using a process called click chemistry. An ordinary GFP molecule does not normally bind with other GFP molecules to form fibers. But the researchers modified the GFP molecules and attached PEO-dialkyne linkers to them, they noticed something strange — the GFP molecules appeared to self-assemble into long fibers.
Importantly, the fibers disassembled after being exposed to sound waves, and then reassembled within a few days. Systems that exhibit this type of reversible fibrous self-assembly have been long sought by scientists for use in applications such as tissue engineering, drug delivery, nanoreactors and imaging.
To determine what processes were driving the self-assembly, Anna Balazs, Distinguished Professor of Chemical Engineering and the Robert v. d. Luft Professor at the University of Pittsburgh ran a computer simulation of the GFP molecules’ self-assembly process using a technique called dissipative particle dynamics, a type of coarse-grained molecular dynamics method.
The simulation confirmed the modified GFP’s tendency to form fibers and revealed that the self-assembly process was driven by the interaction of hydrophobic patches on the surfaces of individual GFP molecules.
“Our protein-polymer system gives us an atomically precise, very well-defined nanoscale building object onto which we can attach different handles in very precisely defined positions. It can be used in a way that wasn’t ever intended by biology,” Kowalewski said.
Researchers at the University of Pittsburgh, Oregon State University, and Zhejiang University also participated in the study. The research was supported by the U.S. Department of Energy, National Science Foundation, Carnegie Mellon’s CRP Consortium and Oregon State University.
Abstract of Angewandte Chemie International Edition paper
We demonstrate a simple bioconjugate polymer system that undergoes reversible self-assembling into extended fibrous structures, reminiscent of those observed in living systems. It is comprised of green fluorescent protein (GFP) molecules linked into linear oligomeric strands through click step growth polymerization with dialkyne poly(ethylene oxide) (PEO). Confocal microscopy, atomic force microscopy, and dynamic light scattering revealed that such strands form high persistence length fibers, with lengths reaching tens of micrometers, and uniform, sub-100 nm widths. We ascribe this remarkable and robust form of self-assembly to the cooperativity arising from the known tendency of GFP molecules to dimerize through localized hydrophobic patches and from their covalent pre-linking with flexible PEO. Dissipative particle dynamics simulations of a coarse-grained model of the system revealed its tendency to form elongated fibrous aggregates, suggesting the general nature of this mode of self-assembly.