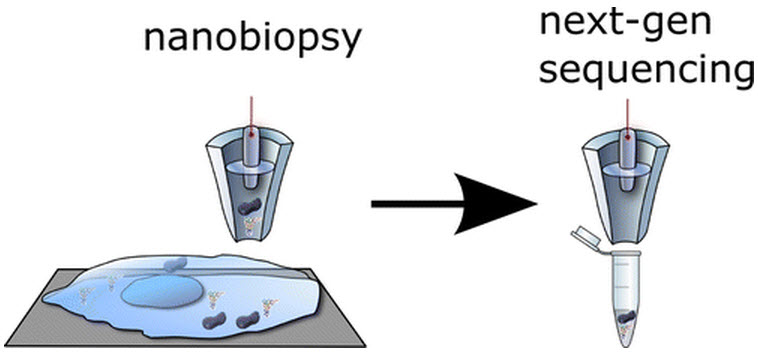
New technique allows minimally invasive ‘nanobiopsies’ of living cells
January 22, 2014
Researchers at UC Santa Cruz (UCSC) have developed a robotic “nanobiopsy” system that can extract tiny samples from inside a living cell without killing it.
The single-cell nanobiopsy technique is a powerful tool for scientists working to understand the dynamic processes that occur within living cells, according to Nader Pourmand, professor of biomolecular engineering in UCSC’s Baskin School of Engineering.
“We can take a biopsy from a living cell and go back to the same cell multiple times over a couple of days without killing it. With other technologies, you have to sacrifice a cell to analyze it,” said Pourmand, who leads the Biosensors and Bioelectrical Technology group at UCSC.
The nanobiopsy platform is the latest device his group has developed that uses nanopipettes, which are small glass tubes that taper to a fine tip with a diameter of just 50 to 100 nanometers. “We can create nanopipettes in the lab — it doesn’t require an expensive nanofabrication facility,” Pourmand said. “To go into a cell, however, the problem is that you cannot see the tip, even with a high-end microscope, so you don’t know how far away from the cell it is.”
Adam Seger, a postdoctoral researcher in the lab (now at MagArray in Sunnyvale), solved this problem by developing a feedback control system based on a customized scanning ion conductance microscope (SICM). The system uses an ion current across the tip of the nanopipette as a feedback signal, detecting a drop in the current when the tip gets close to the cell surface.
An automated control system positions the nanopipette tip just above the cell surface and then plunges it down quickly to penetrate the cell membrane. Manipulating the voltage triggers the controlled uptake of a minute quantity of cellular material.
Minimal cell damage
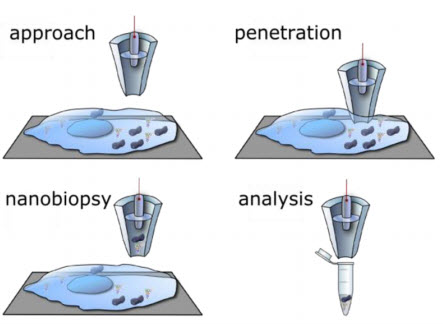
Illustration of automated approach to cell surface, penetration in the cell cytosol, followed by controlled aspiration of cytoplasmic material by electrowetting, and delivery of the biopsied material into a tube for analysis. Scheme not to scale. (Credit: UC Santa Cruz/ACS Nano)
“Because of the small pore of the nanopipette (50 nm in radius), its insertion into cells is minimally invasive. This is a major advance over previous approaches for single cell molecular analysis, which employed micropipets that severely damage cell membranes,” the researchers say in a study published in ACS Nano.
Pourmand’s group used the system to extract from living cells tiny amounts of cellular material estimated to be about 50 femtoliters (a femtoliter is one quadrillionth of a liter).
That’s about one percent of the volume of a human cell. The researchers were able to extract and sequence RNA from individual human cancer cells. They also extracted mitochondria (tiny subcellular organelles) from human fibroblasts and sequenced the mitochondrial DNA.
“Mitochondria are known to be involved in many neurodegenerative diseases. This technology can be used to shed light on the importance of mutations in the mitochondrial genome,” Pourmand said.
There are many potential uses for this technology, and Pourmand said he is eager to develop collaborations with other researchers and explore different applications. “It is a versatile platform for anyone trying to understand what is happening inside the cell, including cancer biologists, stem cell biologists, and others,” he said.
Abstract of ACS Nano paper
The ability to study the molecular biology of living single cells in heterogeneous cell populations is essential for next generation analysis of cellular circuitry and function. Here, we developed a single-cell nanobiopsy platform based on scanning ion conductance microscopy (SICM) for continuous sampling of intracellular content from individual cells. The nanobiopsy platform uses electrowetting within a nanopipette to extract cellular material from living cells with minimal disruption of the cellular milieu. We demonstrate the subcellular resolution of the nanobiopsy platform by isolating small subpopulations of mitochondria from single living cells, and quantify mutant mitochondrial genomes in those single cells with high throughput sequencing technology. These findings may provide the foundation for dynamic subcellular genomic analysis.