Peering deeply into sub-cellular structures
June 5, 2013
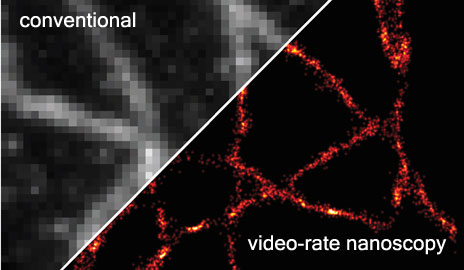
This image of microtubules, which act as a cellular scaffolding, was captured in just 33 milliseconds, or 30 super-resolution images per second (conventional video speed) (credit: Yale University)
A dream of scientists has been to visualize details of two types of important fast-moving sub-cellular cytoskeletal structures — microtubules and actin — within our cells.
Now two groups of researchers have independently succeeded in doing that. Their research could prove vital in the study of cancer and other diseases.
Imaging microtubules in real time
Yale University researchers have developed a way to generate accurate video images of sub-cellular cytoskeletal structures called microtubules in milliseconds, rather than minutes, which are required using the best current microscopes.
F“We can now see research come to life and tackle complex questions or conditions which require hundreds of images, something we have not been able to do before,” said Joerg Bewersdorf, assistant professor of cell biology and biomedical engineering and senior author of the research, published in the journal Nature Methods.
New algorithms
Recently developed cameras using “scientific complementary metal-oxide semiconductors” (sCMOS) — improvements on standard CMOS chips — can dramatically accelerate data acquisition, enlarge the field of view, and increase efficiency. However, the readout noise in these new devices substantially lowers the localization precision and introduces artifacts.
The researchers have developed algorithms that overcome these limitations and provide precise localization of single molecules, with super-resolution imaging at rates of up to 32 reconstructed images per second in both fixed and living cells
Imaging an elusive high-speed biological process
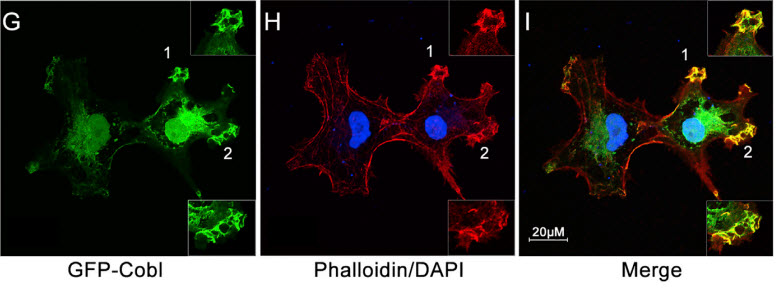
A biological “freeze-frame” showing the initial step in the formation of actin, a sturdy strand-like filament that is vital in all animal and human cells. (G) Cobl (which helps form the actin nucleus), (H) Actin, (I) the two interact in actin nucleation (formation of the nucleus). (Credit: J. Ma/Rice University)
Meanwhile, structural biologists from Rice University and Baylor College of Medicine (BCM) have captured the first three-dimensional crystalline snapshot of another critical but fleeting cellular process — the formation of another cytoskeletal filament, actin, which takes place thousands of times per second in each human cell.
Actin participates in many important cellular processes, including muscle contraction, cell motility, cell division and cytokinesis, vesicle and organelle movement, cell signalling, and the establishment and maintenance of cell junctions and cell shape.
The filaments, which are called F-actin, also play key roles in muscle contraction, cell division and other critical processes. “One of the major distinctions between cancerous cells and healthy cells is their shape,” said study co-author Jianpeng Ma, professor of bioengineering at Rice and the Lodwick T. Bolin Professor of Biochemistry at BCM. “There is a correlation between healthy shape and well-regulated cell growth, and cancer cells are often ugly and ill-shaped compared to healthy cells.”
To capture the actin nucleation (formation), the researchers devised a clever way to stabilize the actin nucleus by preventing further polymerization (forming complex chains of molecules into filaments).
How it was accomplished
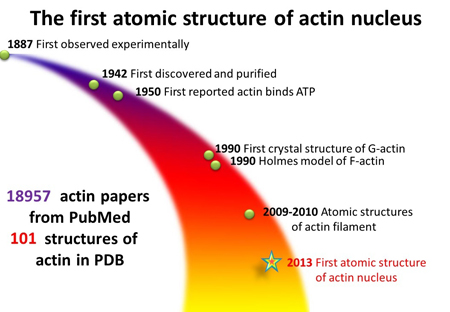 F-actin was discovered in 1887, but despite the more than 18,000 actin-related studies in scientific literature, biologists have struggled to unlock some of its secrets. For example, F-actin is a polymer made of many smaller proteins called monomers.
F-actin was discovered in 1887, but despite the more than 18,000 actin-related studies in scientific literature, biologists have struggled to unlock some of its secrets. For example, F-actin is a polymer made of many smaller proteins called monomers.
These building blocks, which are called G-actin, self-assemble end to end to form F-actin. But the self-assembly process is so efficient that scientists have been unable to see what happens when the first two or three monomers come together to form the nucleus of a filament.
The F-actin filaments inside cells are constantly being built, torn apart and rebuilt. “Nucleation is critical for this continual building and rebuilding,” said BCM biochemist and study co-author Qinghua Wang. “For healthy cells, nucleation is the starting place for robust shape. For unhealthy cells, like cancer, nucleation processes may play a crucial role in unregulated growth. That’s one reason we want to better understand nucleation.”
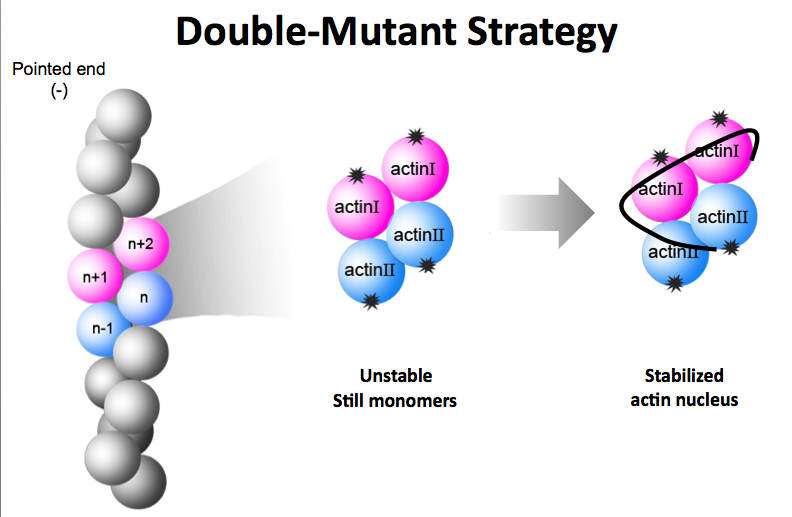
To decipher the structure of the F-actin nucleus, researchers created two mutant versions of actin monomers that could bind together to form a nucleus but could not bind with additional monomers to form the F-actin polymer chain. (Credit: J. Ma/Rice University)
In 2008, Ma and Wang asked Xiaorui Chen, a graduate student in BCM’s Structural and Computational Biology and Molecular Biophysics program, to undertake the task of using x-ray crystallography to determine the structure of the actin nucleus.
Her initial attempts failed, but the team finally hit upon the winning idea of creating two mutant versions of G-actin that could nucleate but not polymerize.
Native G-actin binds with one neighbor on top and one on bottom, and this top-bottom, end-to-end binding pattern is the key to forming long F-actin polymers.
To foster nucleation without polymerization, Chen created two mutant versions of G-actin. One mutant could bind normally on top but not on bottom, and the other could bind normally on bottom but not on top.
“This dual-mutant strategy was the key,” said Chen, who is now a postdoctoral researcher at BCM. “After that, we had to overcome problems related to forming and growing the crystal samples needed for crystallography.”
Chen used a two-stage process to prepare the crystals. She first used high levels of super-saturation to spur initial crystal formation and then used a process called seeding to transfer the newly formed crystals to another medium where they could grow large enough for examination.
Once the crystals were prepared, they were analyzed with x-ray diffraction, which revealed the atomic arrangement of each atom in the nucleated, dual-mutant pair.
“We believe this dual-mutant arrangement reveals the most critical contacts involved in nucleation,” Ma said. “For the first time, we are able to see how actin nucleation begins.”
The research was supported by the National Institutes of Health, the Gillson-Longenbaugh Foundation, the National Science Foundation, the Welch Foundation, the Department of Energy and the Michigan Economic Development Corp.