Researchers reverse bacterial resistance to antibiotics
May 7, 2015
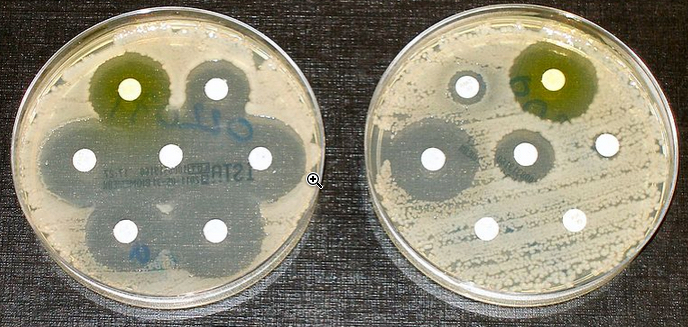
Antibiotic resistance tests: Bacteria in the culture on the left are susceptible to the antibiotic in each disk, as shown by the dark, clear rings where bacteria have not grown. Those on the right are fully susceptible to only three of the seven antibiotics tested. (credit: Graham Beards/Wikimedia Commons)
Biologist Miriam Barlow of the University of California, Merced, and mathematician Kristina Crona of American University of Washington, DC have found a way to return bacteria to a pre-resistant state to help doctors deal with the growing problem of resistant bacteria. In research published in the open-access journal PLOS ONE, they show how to verify treatment options for a family of 15 antibiotics used to fight common infections, including penicillin.
Their work could have major implications for doctors attempting to keep patient infections at bay using “antibiotic cycling,” in which a handful of different antibiotics are used on a rotating basis.
“Doctors don’t take an ordered approach when they rotate antibiotics,” Barlow said. “The doctors would benefit from a system of rotation that is proven. Our goal was to find a precise, ordered schedule of antibiotics that doctors could rely on and know that in the end, resistance will be reversed, and an antibiotic will work.”
Resistance to antibiotics is a natural part of the evolution of bacteria, and unavoidable given the many types of bacteria and the susceptibility of the human host. To compensate for bacterial evolution, a doctor fighting infections in an intensive care unit may reduce, rotate, or discontinue different antibiotics to get them to be effective in the short term.
Strategic antibiotic cycling
The researchers exposed bacteria to 15 different antibiotics and measured their growth rates. From there, they computed the probability of mutations to return the bacteria to its harmless state using “Time Machine” software.
Managing resistance in any drug environment is extremely difficult, because bacteria evolve so quickly, becoming highly resistant after many mutations. To find optimal cycling strategies, the researchers tested up to six drugs in rotation at a time and found optimal plans for reversing the evolution of drug-resistant bacteria.
“This shows antibiotics cycling works. As a medical application, physicians can take a more strategic approach,” Crona said. “Uncovering optimal plans in antibiotics cycling presents a mathematical challenge.”
The researchers hope to next test the treatment paths in a clinical setting, working with doctors to rotate antibiotics to maximize their efficacy.
“This work shows that there is still hope for antibiotics if we use them intelligently,” Barlow said. “More research in this area and more research funding would make it possible to explore the options more comprehensively.”
Abstract of Rational Design of Antibiotic Treatment Plans: A Treatment Strategy for Managing Evolution and Reversing Resistance
The development of reliable methods for restoring susceptibility after antibiotic resistance arises has proven elusive. A greater understanding of the relationship between antibiotic administration and the evolution of resistance is key to overcoming this challenge. Here we present a data-driven mathematical approach for developing antibiotic treatment plans that can reverse the evolution of antibiotic resistance determinants. We have generated adaptive landscapes for 16 genotypes of the TEM β-lactamase that vary from the wild type genotype “TEM-1” through all combinations of four amino acid substitutions. We determined the growth rate of each genotype when treated with each of 15 β-lactam antibiotics. By using growth rates as a measure of fitness, we computed the probability of each amino acid substitution in each β-lactam treatment using two different models named the Correlated Probability Model (CPM) and the Equal Probability Model (EPM). We then performed an exhaustive search through the 15 treatments for substitution paths leading from each of the 16 genotypes back to the wild type TEM-1. We identified optimized treatment paths that returned the highest probabilities of selecting for reversions of amino acid substitutions and returning TEM to the wild type state. For the CPM model, the optimized probabilities ranged between 0.6 and 1.0. For the EPM model, the optimized probabilities ranged between 0.38 and 1.0. For cyclical CPM treatment plans in which the starting and ending genotype was the wild type, the probabilities were between 0.62 and 0.7. Overall this study shows that there is promise for reversing the evolution of resistance through antibiotic treatment plans.