A DNA-based nanosensor that detects cancer by its pH
May 13, 2014
Cartoon showing a DNA-based nanosensor that allows for measuring pH variation at the nanoscale for used in cancer in-vivo imaging and targeted drug-delivery. This nanosensor measures less than 10 nanometers and unfolds at a specifically programmed pH. (Credit: Marco Tripodi)
Bioengineers at the University of Rome, Tor Vergata and the University of Montreal have used DNA to develop a tool that detects and reacts to chemical changes caused by cancer cells. It may one day be used to deliver drugs to tumor cells.
The researchers’ nanosensor measures pH variations at the nanoscale, indicating how acidic (a lower pH level) or alkaline (a higher pH level).
“In living organisms, these small pH changes typically occur in tiny areas measuring only a few hundred nanometers,” says senior author Prof. Francesco Ricci. “Developing sensors or nanomachines that can measure pH changes at this scale should prove of utility for several applications in the fields of in-vivo imaging, clinical diagnostics, and drug-delivery.”
“DNA represents an ideal material to build sensors or nanomachines at the nanometer scale” says senior author Prof. Vallée-Bélisle. “By taking advantage of a specific DNA sequences that form pH-sensitive triple helix, we have designed a versatile nanosensor that can be programmed to fluoresce [shine] only at specific pH values” that are characteristic of a specific disease.
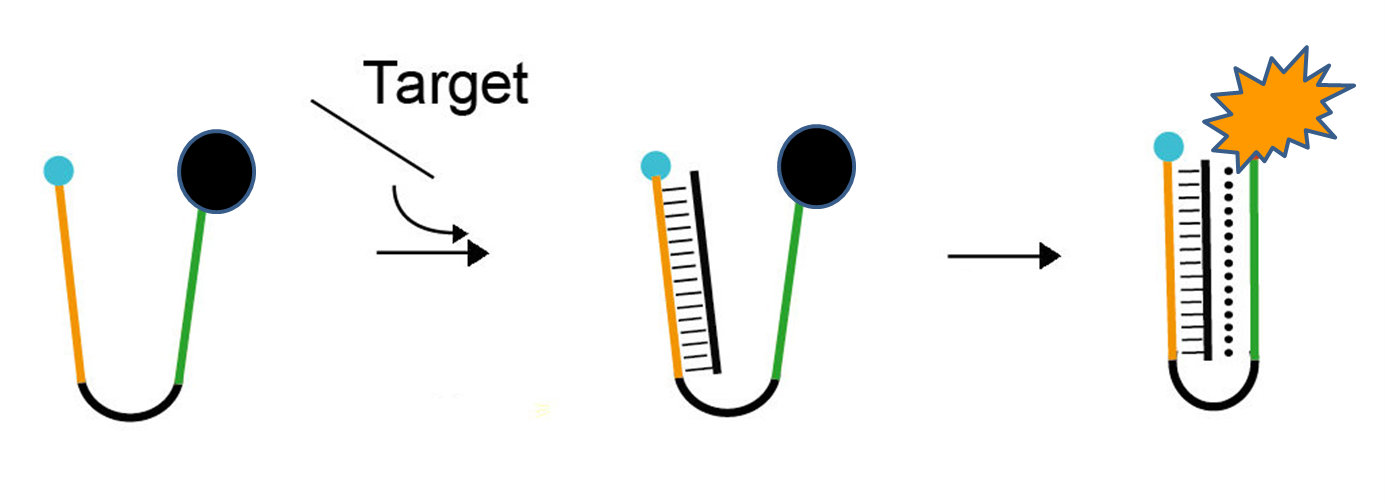
A pH-sensitive nanoswitch binds to the DNA target, resulting in the clamp closing and activating the fluorescent signal (orange). (Credit: Francesco Ricci, University of Rome, Tor Vergata)
For example, cancer cells often display a lower pH compared to normal cells (the pH level inside cancer cells is higher than it is outside).
So in the future, this recently patented nanotechnology may also find applications in the development of novel drug-delivery platforms that release drugs only in the vicinity of tumor cells.
This research was published in the Journal of American Chemical Society and was supported by the European Research Council and the Natural Sciences and Engineering Research Council of Canada.
UPDATE 5/13/2014: pH definition corrected
Abstract of Journal of American Chemical Society paper
We have designed programmable DNA-based nanoswitches whose closing/opening can be triggered over specific different pH windows. These nanoswitches form an intramolecular triplex DNA structure through pH-sensitive parallel Hoogsteen interactions. We demonstrate that by simply changing the relative content of TAT/CGC triplets in the switches, we can rationally tune their pH dependence over more than 5 pH units. The ability to design DNA-based switches with tunable pH dependence provides the opportunity to engineer pH nanosensors with unprecedented wide sensitivity to pH changes. For example, by mixing in the same solution three switches with different pH sensitivity, we developed a pH nanosensor that can precisely monitor pH variations over 5.5 units of pH. With their fast response time (<200 ms) and high reversibility, these pH-triggered nanoswitches appear particularly suitable for applications ranging from the real-time monitoring of pH changes in vivo to the development of pH sensitive smart nanomaterials.
Abstract of ACS Nano paper
Naturally occurring chemoreceptors almost invariably employ structure-switching mechanisms, an observation that has inspired the use of biomolecular switches in a wide range of artificial technologies in the areas of diagnostics, imaging, and synthetic biology. In one mechanism for generating such behavior, clamp-based switching, binding occurs via the clamplike embrace of two recognition elements onto a single target molecule. In addition to coupling recognition with a large conformational change, this mechanism offers a second advantage: it improves both affinity and specificity simultaneously. To explore the physics of such switches we have dissected here the thermodynamics of a clamp-switch that recognizes a target DNA sequence through both Watson-Crick base pairing and triplex-forming Hoogsteen interactions. When compared to the equivalent linear DNA probe (which relies solely on Watson-Crick interactions), the extra Hoogsteen interactions in the DNA clamp-switch increase the probe’s affinity for its target by  0.29 ± 0.02 kcal/mol/base. The Hoogsteen interactions of the clamp-switch likewise provide an additional specificity check that increases the discrimination efficiency toward a single-base mismatch by 1.2 ± 0.2 kcal/mol. This, in turn, leads to a 10-fold improvement in the width of the “specificity window” of this probe relative to that of the equivalent linear probe. Given these attributes, clamp-switches should be of utility not only for sensing applications but also, in the specific field of DNA nanotechnology, for applications calling for a better control over the building of nanostructures and nanomachines.
0.29 ± 0.02 kcal/mol/base. The Hoogsteen interactions of the clamp-switch likewise provide an additional specificity check that increases the discrimination efficiency toward a single-base mismatch by 1.2 ± 0.2 kcal/mol. This, in turn, leads to a 10-fold improvement in the width of the “specificity window” of this probe relative to that of the equivalent linear probe. Given these attributes, clamp-switches should be of utility not only for sensing applications but also, in the specific field of DNA nanotechnology, for applications calling for a better control over the building of nanostructures and nanomachines.