Rewriting DNA to understand its ‘language’ and improve gene therapy
June 1, 2012
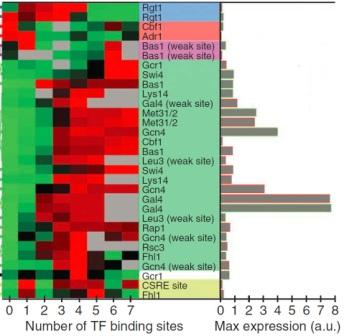
The relationship between gene activity and the number of binding sites for regulatory proteins in the gene’s control region. The red-green scale shows gene activity levels (red is high). Gray bars show the maximal level of gene activity achieved by each type of regulatory protein. (Credit: Prof. Eran Segal/Weizmann Institute)
New Weizmann Institute technology speeds up DNA “rewriting” (altering) and measures the effects of the changes in living cells.
To advance our understanding of the genetic code, a new Weizmann Institute study proposes a way of introduce DNA segments into genomes of living cells and test the effects of these changes.
Until now, changing the DNA sequence has been a slow and labor-intensive process. It took several weeks to alter just one DNA region at a time; testing the effects of each of these changes took even longer.
Understanding the hidden DNA language
Weizmann Institute scientists have developed a technology that makes it possible to simultaneously introduce tens of thousands of DNA regions into tens of thousands of living cells — each region in a separate cell — in a planned and systematic manner, and measure the results of each such change with great precision and in a single experiment.
“This fast method will significantly advance our ability to understand the ‘language’ of DNA,” says research team leader Prof. Eran Segal, of the Weizmann Institute’s Computer Science and Applied Mathematics and Molecular Cell Biology Departments. “Reading out a person’s entire genome is already a manageable task, but what exactly is written in that genome?”
Why we need to identify DNA “words” and understand their meaning
- Understand how genotypic differences among people generate observable differences among them — from the way we look to the way our cells function. For example, which genetic differences are responsible for the development of various diseases in certain individuals?
- Improve genetic therapies based on introducing new genes or improve regulatory sequences into cells to repair genetic defects.
- Understand how the control of gene expression is encoded in the DNA — that is, the instructions determining the level of activity of each gene in the genetic code — one of the central in molecular biology, because gene activity levels have crucial effects on cell function. For example, how a gene’s activity level is affected by the gene’s distance from its regulatory sequence.
The new method consists of four steps: (1) creation of 50,000 different genetic sequences on DNA chips, (2) massive insertion of these sequences into cells at the same time, (3) sorting the cells with the help of a sorting machine that senses the expression levels of a “reporter” gene, and (4) high-throughput parallel DNA sequencing.
Ref.: Eilon Sharon et al., Inferring gene regulatory logic from high-throughput measurements of thousands of systematically designed promoters, Nature Biotechnology, 2012, DOI: 10.1038/nbt.2205
Ref.: Tali Raveh-Sadka et al., Manipulating nucleosome disfavoring sequences allows fine-tune regulation of gene expression in yeast, Nature Genetics, 2012, DOI: 10.1038/ng.2305