Woman dies from antibiotic-resistant bacteria when no antibiotics worked
January 18, 2017
Carbapenem-resistant Enterobacteriaceae (CRE) bacteria (credit: Melissa Brower/CDC)
The death of a hospitalized patient in Reno Nevada for whom no available antibiotics worked highlights what World Health Organization and other public-health experts have been warning: antibiotic resistance is a serious threat and has gone global.
The patient — a female in her 70s — was admitted in for an infection and died in September 2016 from septic shock the CDC announced on Jan. 13. The patient had been treated for multiple infections in India before traveling to the United States. The infection that led to her hospitalization in Reno was caused by a strain of carbapenem-resistant Enterobacteriaceae (CRE)* bacteria known as Klebsiella pneumoniae. Although not all strains of Klebsiella pneumonia are CRE, the strain that infected this patient was resistant to all available antibiotics, according to the CDC. (Carbapeneum is a “drug of last resort.”)
In a paper in The Lancet in October, researchers reported that more than a third of blood infections in newborn babies involving Klebsiella pneumoniae and similar bacteria were resistant to multiple drugs to the point they were virtually untreatable and “threaten the return of a pre-antibiotic era in Indian neonatal intensive care units,” the study’s authors warned.
Spreading more widely and stealthily that thought
Highlighting the importance of this report, a new NIH-funded study published online January 16, 2017 in PNAS (Proceedings of the National Academy of Sciences) from Harvard T.H. Chan School of Public Health and the Broad Institute of MIT and Harvard found that the CRE family of highly drug-resistant and potentially deadly bacteria may be spreading more widely — and more stealthily — than previously thought.
The researchers examined CRE bacteria that were isolated from patients with infections at four U.S. hospitals. They found a wide variety of CRE . They also found a diverse array of genetic traits enabling CRE to resist antibiotics, and found that these traits are transferring easily among various CRE species.
The findings suggest that CRE may well be transmitting from person to person asymptomatically, and that genomic surveillance of this dangerous bacteria should be increased.
CRE are a class of bacteria that is resistant to multiple antibiotics, including carbapenems, which are considered last-resort drugs when other antibiotics have failed. CRE, which tend to spread in hospitals and long-term care facilities, cause an estimated 9,300 infections and 600 deaths in the U.S. each year, according to the CDC — and incidence is on the rise. CDC director Tom Frieden has called these “nightmare bacteria” because they are resistant to some of the last-ditch treatments available to doctors battling resistant infections.
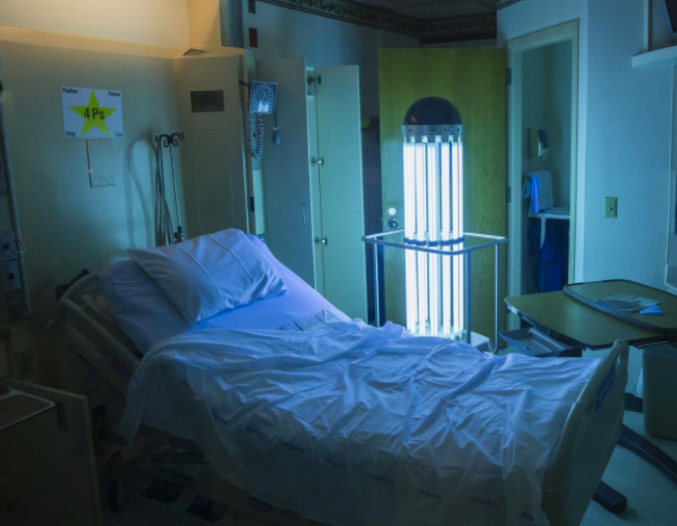

A type of ultraviolet light called UVC could aid hospitals in the ongoing battle to keep drug-resistant bacteria from lingering in patient rooms and causing new infections. A CDC-NIH-funded study led by Duke Health and published in The Lancet finds use of UVC machines can cut transmission of four major superbugs by a cumulative 30 percent. (credit: Shawn Rocco/Duke Health)
Unobserved transmissions
“While the typical focus has been on treating sick patients with CRE-related infections, our new findings suggest that CRE is spreading beyond the obvious cases of disease,” said William Hanage, associate professor of epidemiology at Harvard Chan School and senior author of the study. “We need to look harder for this unobserved transmission within our communities and healthcare facilities if we want to stamp it out.”
The researchers looked at about 250 samples of CRE from hospitalized patients from three Boston-area hospitals and from one California hospital.
The researchers found what Hanage termed a “riot of diversity,” both among CRE species and among carbapenem resistance genes. They also found that resistance genes are moving easily from species to species, contributing to a continually evolving threat from CRE.
In addition, the researchers found that some CRE bacteria employ uncharacterized resistance mechanisms—implying that there are more to be discovered. The finding highlights the need for vigilance in searching for as yet unknown forms of resistance as they evolve and emerge.
What if there were no new antibacterial drugs?
But what’s also needed is new antibiotics. “Meaningful incentives for pharmaceutical companies have been suggested to encourage them to re-enter this area of drug development; these incentives include models to push companies to develop these drugs or by making it economically viable so as to pull companies back to this area as well as alternative routes of funding,” notes Prof Laura J V Piddock, Antimicrobial Agents Research Group, School of Immunity and Infection, College of Medical and Dental Sciences, University of Birmingham in “The crisis of no new antibiotics—what is the way forward?” in Lancet Infect Dis. in 2012.
“Also, few data exist on the human and economic costs of antibacterial resistance or the potential costs of inaction (i.e., what would the social effects be if no new antibacterial drugs were available and fatal infections became the norm?). To assess such costs, involvement of governmental agencies that recommend use of drugs is needed (i.e., FDA, European Medicines Agency, and in individual countries such as UK, The National Institute for Health and Clinical Excellence).”
* “Carbapenems are one of few antimicrobials that have been effective against multidrug-resistant bacteria, but their utility is threatened by the emergence of carbapenem-resistant Enterobacteriaceae (CRE),” according to an open-access PLoS ONE paper published in 2014. “Klebsiella pneumoniae is the most common CRE species in the United States, typically encountered as a hospital-acquired infection with high morbidity and mortality, and resistant to nearly all available antibiotics.” In this case, metallo-β-lactamase NDM-1 (New Delhi metallo-beta-lactamase or NDM), a version of Klebsiella pneumoniae, was found.