Robot biologist solves complex problem from scratch
October 17, 2011
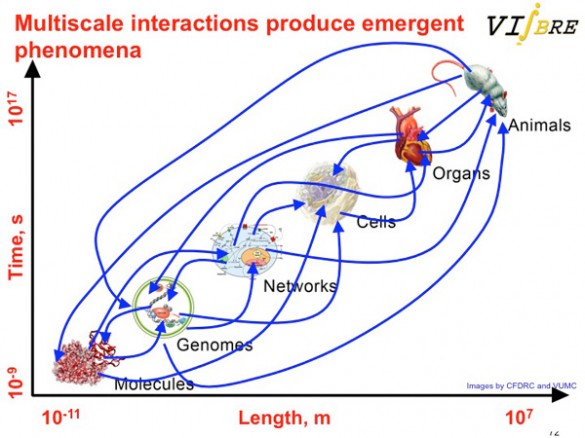
One of the things that makes biology so complicated is that processes at different scales ranging from the molecular to whole animals are continually interacting with each other (credit: Wikswo Lab)
An interdisciplinary team of scientists has taken a major step toward automating the scientific process with the Automated Biology Explorer (ABE) system, which can analyze raw experimental data from a biological system and derive the basic mathematical equations that describe the way the system operates.
According to the researchers at Vanderbilt University, Cornell University and CFD Research Corporation, it is one of the most complex scientific modeling problems that a computer has solved completely from scratch.
The work was a collaboration between John P. Wikswo, the Gordon A. Cain University Professor at Vanderbilt, Michael Schmidt and Hod Lipson at the Creative Machines Lab at Cornell University and Jerry Jenkins and Ravishankar Vallabhajosyula at CFDRC in Huntsville, Ala.
John P. Wikswo, the Gordon A. Cain University Professor at Vanderbilt, has christened the is a unique piece of software called
ABE’s “brain” is software called Eureqa, developed at Cornell in 2009. One of Eureqa’s initial achievements was identifying the basic laws of motion by analyzing the motion of a double pendulum. What took Sir Isaac Newton years to discover, Eureqa did in a few hours when running on a personal computer.
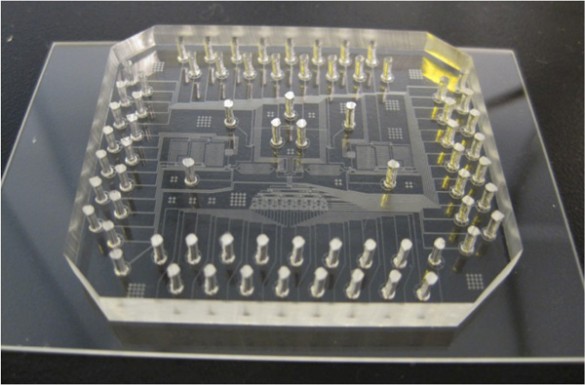

A microformulator that will give ABE the ability to perform experiments without human intervention (credit: Wikswo Lab)
Software derives biochemical equations automatically
The biological system that the researchers used to test ABE is glycolysis, the primary process that produces energy in a living cell. They focused on how yeast cells control glycolytic oscillations because it is one of the most extensively studied biological control systems. ABE derived the equations a priori. The only thing the software knew in advance was addition, subtraction, multiplication and division.
The ability to generate mathematical equations from scratch is what sets ABE apart from Adam, the robot scientist developed by Ross King and his colleagues at the University of Wales at Aberystwyth. Adam runs yeast genetics experiments and made international headlines two years ago by making a novel scientific discovery without direct human input. King fed Adam with a model of yeast metabolism and a database of genes and proteins involved in metabolism in other species. He also linked the computer to a remote-controlled genetics laboratory. This allowed the computer to generate hypotheses, then design and conduct actual experiments to test them.
To give ABE the ability to run experiments like Adam, tbe researchers are urrently developing “laboratory-on-a-chip” technology that can be controlled by Eureqa. This will allow ABE to design and perform a wide variety of basic biology experiments. Their initial effort is focused on developing a microfluidics device that can test cell metabolism.
Why biology needs automation
Biology is more complex than astronomy or physics or chemistry,” maintained John P. Wikswo, the Gordon A. Cain University Professor at Vanderbilt. “In fact, it may be too complex for the human brain to comprehend.”
This complexity stems from the fact that biological processes range in size from the dimensions of an atom to those of a whale and in time from a billionth of a second to billions of seconds. Biological processes also have a tremendous dynamic range: for example, the human eye can detect a star at night that is one billionth as bright as objects viewed on a sunny day.
Then there is the matter of sheer numbers. A cell expresses between 10,000 to 15,000 proteins at any one time. Proteins perform all the basic tasks in the cell, including producing energy, maintaining cell structures, regulating these processes and serving as signals to other cells. At any one time, there can be anywhere from three to 10 million copies of a given protein in the cell.
According to Wikswo, the crowning source of complication is that processes at all these different scales interact with one another: “These multi-scale interactions produce emergent phenomena, including life and consciousness.”
Looked at from a mathematical point of view, to create an accurate model of a single mammalian cell may require generating and then solving somewhere between 100,000 to one million equations.
Balanced against this complexity is the capability of the human brain. The biophysicist cites research that has found that the human brain can only process seven pieces of data at a time and quotes a 1938 assessment of brain research by Emerson Pugh: “If the human brain were so simple that we could understand it, we would be so simple that we couldn’t.”
That is where robot scientists like ABE and Adam come in, Wikswo argues. They have the potential for both generating and analyzing the tremendous amounts of data required to really understand how biological systems work and predict how they will react to different conditions.
Co-evolution
“We set out to work with robots, but our path took us, through many twists and turns, to automating science,” said Hod Lipson at the Creative Machines Lab at Cornell University.
His starting point was an attempt to breed robot control systems using an approach modeled on natural selection, instead of having a programmer code in all the steps. Individual programming had largely broken down as robots became more complex because the robots didn’t perform correctly without extensive and time-consuming debugging.
Lipson used genetic programming for the breeding process. It involves starting with the basic components of a robot, randomly combining them in millions of different configurations and then testing how well they perform by a specific criterion, such as how fast they can move. The designs that work the best are then randomly combined and tested. These steps are repeated until it produces a design that is acceptable. However, this process also proved to be too slow.
So Lipson combined the breeding and the debugging processes in an approach he calls co-evolution. He started with a crude simulator, used it to design a robot, tested the design, and studied how it failed. He used this information to improve the simulator so that it could predict the failure. Then he used the improved simulator to design another robot, tested the design, watched how it failed and improved the simulator once again. Repeating these steps of co-evolving simulators and robots produced increasingly competent designs, he found.
After proving that co-evolution works for robot design, Lipson realized that it could be generalized to solve other problems. Specifically, he adapted it for the mathematical process of curve fitting, more generally called symbolic regression. This involves deriving equations that can describe various data sets. Lipson’s software package, Eureqa, proved to be extremely successful. As the word got around, he began getting requests for copies of the program and decided to make it into a citizen science project, available for anyone to download on the Internet.
“Today, it has more than 20,000 users. People are using it to solve problems in a wide variety of areas including traffic, business and neighborhood problems,” Lipson said. Wikswo says this approach will give scientists the ability to control biological systems even if they can’t completely explain how they work, and will allow for developing significantly improved drugs and other therapies.
Ref.: Michael D Schmidt, et al., Automated refinement and inference of analytical models for metabolic networks, Physical Biology, 2011; 8 (5): 055011 [DOI: 10.1088/1478-3975/8/5/055011]